Probe CUST_6602_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6602_PI426222305 | JHI_St_60k_v1 | DMT400036705 | CACAAGAGTCATATATGATACAAAACAATCACAAGTTGGATTTGCAAAAGAAACCTGCAG |
All Microarray Probes Designed to Gene DMG400014156
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6602_PI426222305 | JHI_St_60k_v1 | DMT400036705 | CACAAGAGTCATATATGATACAAAACAATCACAAGTTGGATTTGCAAAAGAAACCTGCAG |
| CUST_6668_PI426222305 | JHI_St_60k_v1 | DMT400036706 | GCCAATAACGACTCGTTGGCGATAATATCTTTGCTTAACATACATAACTATTTGAGGTTT |
Microarray Signals from CUST_6602_PI426222305
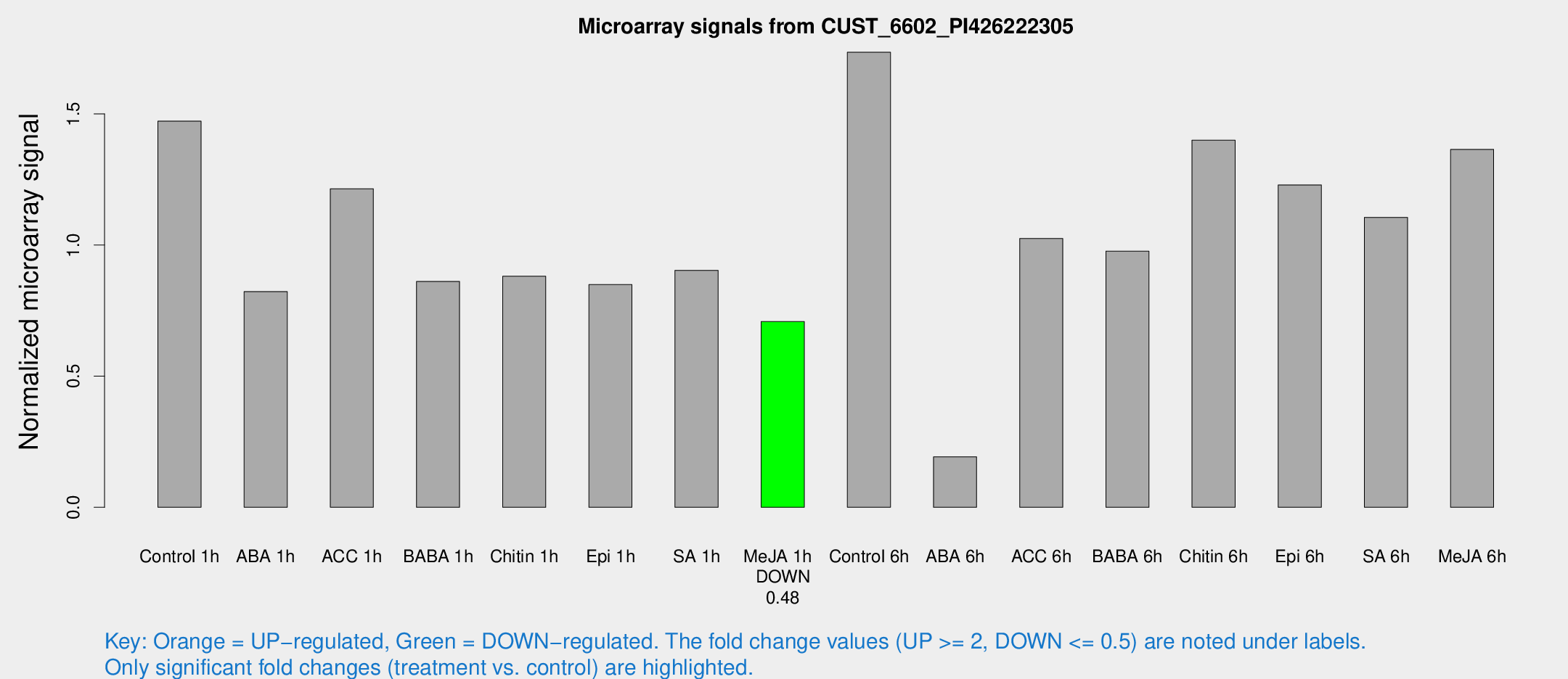
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 842.743 | 93.3105 | 1.47288 | 0.0852943 |
| ABA 1h | 418.107 | 52.2965 | 0.822769 | 0.0948934 |
| ACC 1h | 716.868 | 89.9282 | 1.21474 | 0.0706216 |
| BABA 1h | 502.821 | 118.335 | 0.861386 | 0.150639 |
| Chitin 1h | 460.97 | 73.2203 | 0.88105 | 0.0964537 |
| Epi 1h | 420.202 | 42.9713 | 0.849177 | 0.0810353 |
| SA 1h | 531.529 | 58.1858 | 0.903713 | 0.0729062 |
| Me-JA 1h | 329.727 | 29.4707 | 0.70843 | 0.0417265 |
| Control 6h | 1034.98 | 212.226 | 1.73544 | 0.248502 |
| ABA 6h | 120.362 | 21.8103 | 0.192624 | 0.0419783 |
| ACC 6h | 687.373 | 116.948 | 1.02508 | 0.137935 |
| BABA 6h | 631.758 | 96.6562 | 0.976746 | 0.149131 |
| Chitin 6h | 844.735 | 49.0958 | 1.39986 | 0.113031 |
| Epi 6h | 816.884 | 152.356 | 1.2295 | 0.364266 |
| SA 6h | 639.335 | 108.793 | 1.10571 | 0.0802921 |
| Me-JA 6h | 801.863 | 153.425 | 1.36481 | 0.183702 |
Source Transcript PGSC0003DMT400036705 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G79720.1 | +1 | 0.0 | 528 | 263/422 (62%) | Eukaryotic aspartyl protease family protein | chr1:29997259-29998951 REVERSE LENGTH=484 |
