Probe CUST_6420_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6420_PI426222305 | JHI_St_60k_v1 | DMT400014687 | AGACTATTTGGTGGAGTGTCTAATGCAATATCTACTGACATTACAATGGCGGCAGCGACG |
All Microarray Probes Designed to Gene DMG400005732
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6350_PI426222305 | JHI_St_60k_v1 | DMT400014688 | AAAGTTTCTAACTTTACCCCCAATTTCAAGAACCCCAAAGTCGTTACAGCCTGCATGGCG |
| CUST_6420_PI426222305 | JHI_St_60k_v1 | DMT400014687 | AGACTATTTGGTGGAGTGTCTAATGCAATATCTACTGACATTACAATGGCGGCAGCGACG |
Microarray Signals from CUST_6420_PI426222305
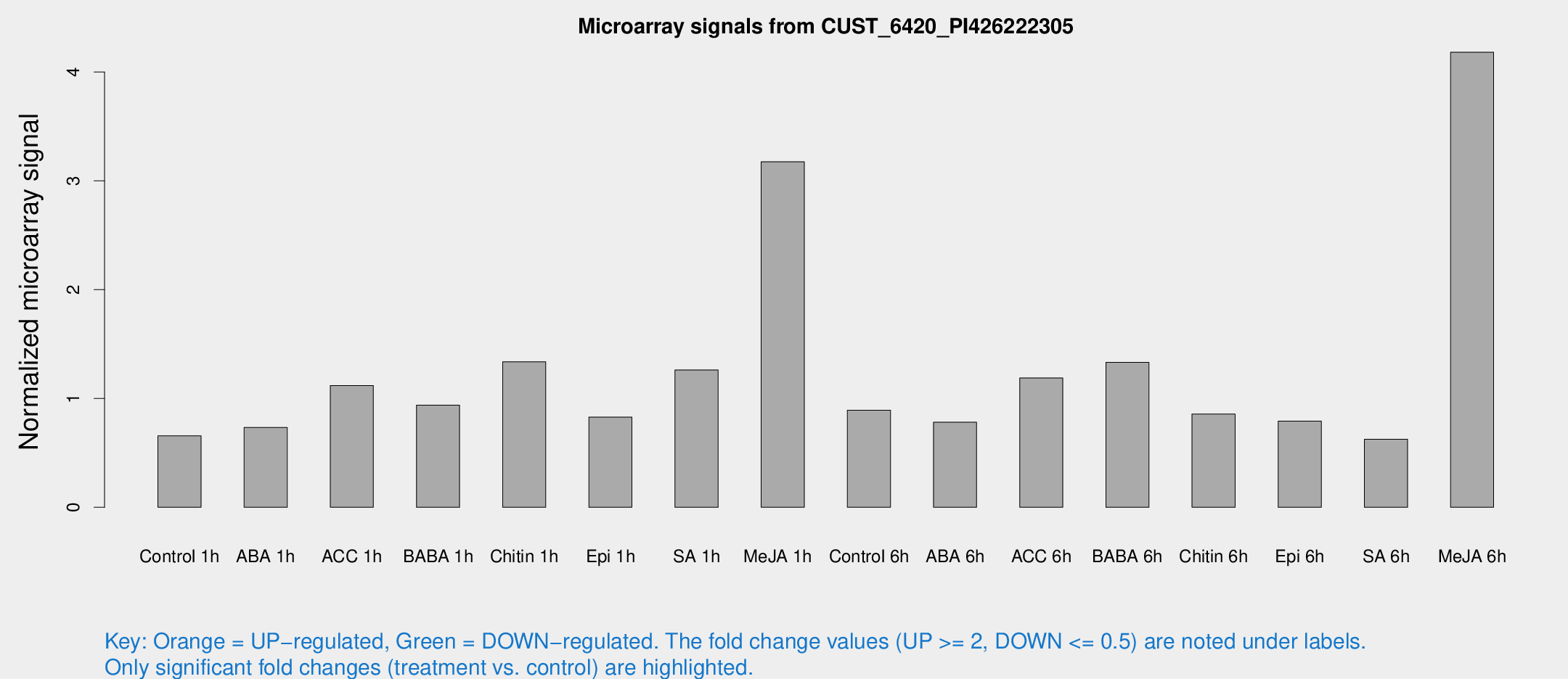
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25.1476 | 5.17599 | 0.656365 | 0.12696 |
| ABA 1h | 25.6149 | 7.5 | 0.734149 | 0.262045 |
| ACC 1h | 44.1751 | 10.6693 | 1.11941 | 0.371184 |
| BABA 1h | 33.608 | 4.58337 | 0.938721 | 0.105315 |
| Chitin 1h | 47.1827 | 11.4016 | 1.33794 | 0.319618 |
| Epi 1h | 27.5336 | 6.5626 | 0.828601 | 0.216635 |
| SA 1h | 48.6451 | 8.13958 | 1.26207 | 0.182952 |
| Me-JA 1h | 106.302 | 37.84 | 3.17493 | 1.1242 |
| Control 6h | 41.9292 | 16.2116 | 0.891788 | 0.502675 |
| ABA 6h | 32.26 | 8.33148 | 0.781825 | 0.176233 |
| ACC 6h | 55.5824 | 19.6122 | 1.18826 | 0.23153 |
| BABA 6h | 56.5943 | 12.1034 | 1.33246 | 0.230991 |
| Chitin 6h | 33.6245 | 3.826 | 0.857 | 0.0999648 |
| Epi 6h | 32.9489 | 3.93445 | 0.792444 | 0.0953775 |
| SA 6h | 30.8936 | 12.7743 | 0.625221 | 0.414859 |
| Me-JA 6h | 154.565 | 21.091 | 4.18192 | 0.981887 |
Source Transcript PGSC0003DMT400014687 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G18270.1 | +1 | 2e-177 | 507 | 261/373 (70%) | cytochrome P450, family 77, subfamily A, polypeptide 5 pseudogene | chr3:6262010-6264025 FORWARD LENGTH=410 |
