Probe CUST_6350_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6350_PI426222305 | JHI_St_60k_v1 | DMT400014688 | AAAGTTTCTAACTTTACCCCCAATTTCAAGAACCCCAAAGTCGTTACAGCCTGCATGGCG |
All Microarray Probes Designed to Gene DMG400005732
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_6350_PI426222305 | JHI_St_60k_v1 | DMT400014688 | AAAGTTTCTAACTTTACCCCCAATTTCAAGAACCCCAAAGTCGTTACAGCCTGCATGGCG |
| CUST_6420_PI426222305 | JHI_St_60k_v1 | DMT400014687 | AGACTATTTGGTGGAGTGTCTAATGCAATATCTACTGACATTACAATGGCGGCAGCGACG |
Microarray Signals from CUST_6350_PI426222305
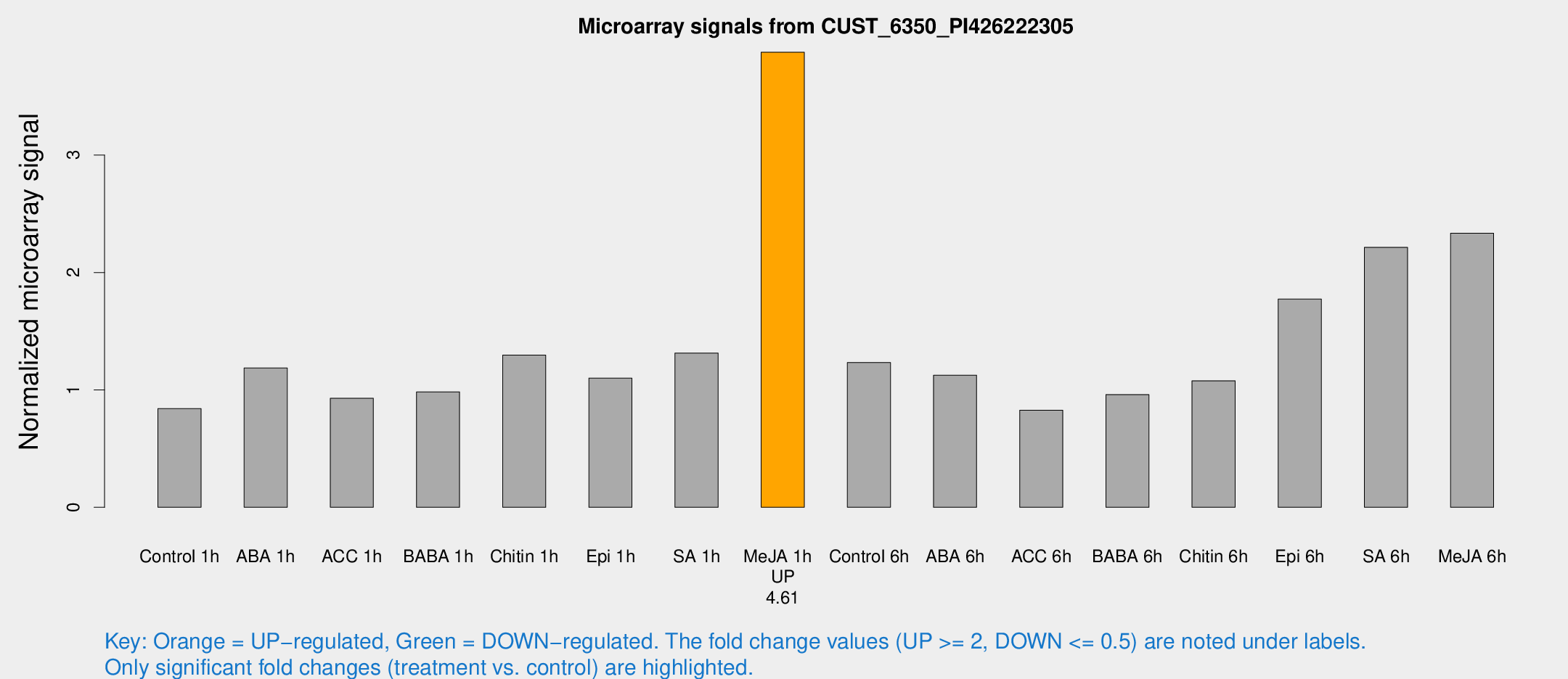
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.23561 | 3.44411 | 0.840625 | 0.462938 |
| ABA 1h | 8.57918 | 3.30548 | 1.18671 | 0.56069 |
| ACC 1h | 7.19851 | 4.24974 | 0.928647 | 0.537978 |
| BABA 1h | 7.2031 | 3.74666 | 0.983127 | 0.525135 |
| Chitin 1h | 9.05452 | 3.45876 | 1.29663 | 0.562615 |
| Epi 1h | 7.58427 | 3.35161 | 1.10005 | 0.543688 |
| SA 1h | 10.8223 | 3.45349 | 1.31363 | 0.501253 |
| Me-JA 1h | 23.9008 | 3.7872 | 3.87603 | 0.619573 |
| Control 6h | 10.3785 | 3.85762 | 1.23334 | 0.574989 |
| ABA 6h | 9.16097 | 3.76611 | 1.12533 | 0.492984 |
| ACC 6h | 7.20184 | 4.29116 | 0.827101 | 0.480551 |
| BABA 6h | 8.01588 | 3.99743 | 0.959984 | 0.480129 |
| Chitin 6h | 8.79941 | 3.99968 | 1.07789 | 0.531336 |
| Epi 6h | 25.9808 | 18.6809 | 1.77366 | 1.96886 |
| SA 6h | 38.8091 | 32.3812 | 2.21425 | 3.82133 |
| Me-JA 6h | 18.1981 | 4.30382 | 2.33392 | 0.537548 |
Source Transcript PGSC0003DMT400014688 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G18270.1 | +2 | 1e-175 | 508 | 270/409 (66%) | cytochrome P450, family 77, subfamily A, polypeptide 5 pseudogene | chr3:6262010-6264025 FORWARD LENGTH=410 |
