Probe CUST_5396_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5396_PI426222305 | JHI_St_60k_v1 | DMT400009095 | CGTCGCTCTAGATCGCTTGCTGGTATGAGATCGCTTAACTTATATGGATTGACAGTATAA |
All Microarray Probes Designed to Gene DMG400003536
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5352_PI426222305 | JHI_St_60k_v1 | DMT400009094 | CAAGCAAAACTAACTGTGTTCTTCATTATTTACATCCTTAATCTCTCCAACATGTTGCTG |
| CUST_5396_PI426222305 | JHI_St_60k_v1 | DMT400009095 | CGTCGCTCTAGATCGCTTGCTGGTATGAGATCGCTTAACTTATATGGATTGACAGTATAA |
Microarray Signals from CUST_5396_PI426222305
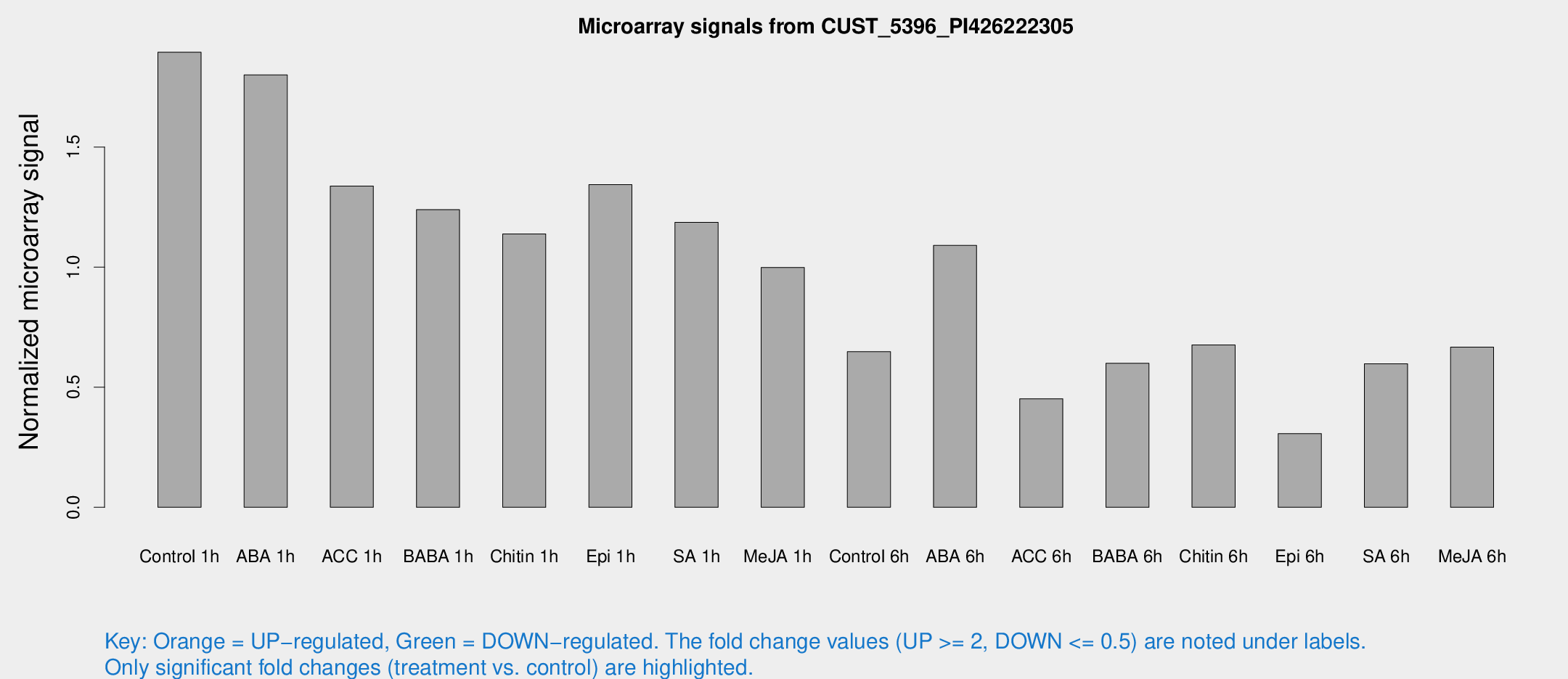
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 151.291 | 15.2063 | 1.89448 | 0.117925 |
| ABA 1h | 126.315 | 8.02379 | 1.80024 | 0.206123 |
| ACC 1h | 127.211 | 42.6216 | 1.33716 | 0.516128 |
| BABA 1h | 98.4829 | 19.6628 | 1.23886 | 0.149629 |
| Chitin 1h | 81.9014 | 9.00648 | 1.13778 | 0.0829538 |
| Epi 1h | 94.7724 | 16.6507 | 1.34299 | 0.202077 |
| SA 1h | 97.2882 | 8.9836 | 1.18659 | 0.154893 |
| Me-JA 1h | 64.6145 | 5.1658 | 0.998533 | 0.0797382 |
| Control 6h | 57.518 | 17.7289 | 0.647973 | 0.185035 |
| ABA 6h | 91.8175 | 6.45859 | 1.09019 | 0.0765783 |
| ACC 6h | 44.1716 | 11.0332 | 0.452086 | 0.0875981 |
| BABA 6h | 53.429 | 5.04634 | 0.599531 | 0.0566477 |
| Chitin 6h | 57.3896 | 5.48173 | 0.675666 | 0.0640787 |
| Epi 6h | 27.6247 | 4.39811 | 0.306476 | 0.0486716 |
| SA 6h | 49.8747 | 13.3279 | 0.597508 | 0.135157 |
| Me-JA 6h | 53.4232 | 7.006 | 0.666655 | 0.0597723 |
Source Transcript PGSC0003DMT400009095 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G36380.1 | +1 | 3e-107 | 328 | 183/291 (63%) | Cytochrome P450 superfamily protein | chr4:17187973-17192202 REVERSE LENGTH=524 |
