Probe CUST_5385_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5385_PI426222305 | JHI_St_60k_v1 | DMT400003959 | CGTATGAGGTTATTGCTGATTATGAAAAACAGGGTCTTGATCGACTCCAGATTATGCCTG |
All Microarray Probes Designed to Gene DMG400001560
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5385_PI426222305 | JHI_St_60k_v1 | DMT400003959 | CGTATGAGGTTATTGCTGATTATGAAAAACAGGGTCTTGATCGACTCCAGATTATGCCTG |
Microarray Signals from CUST_5385_PI426222305
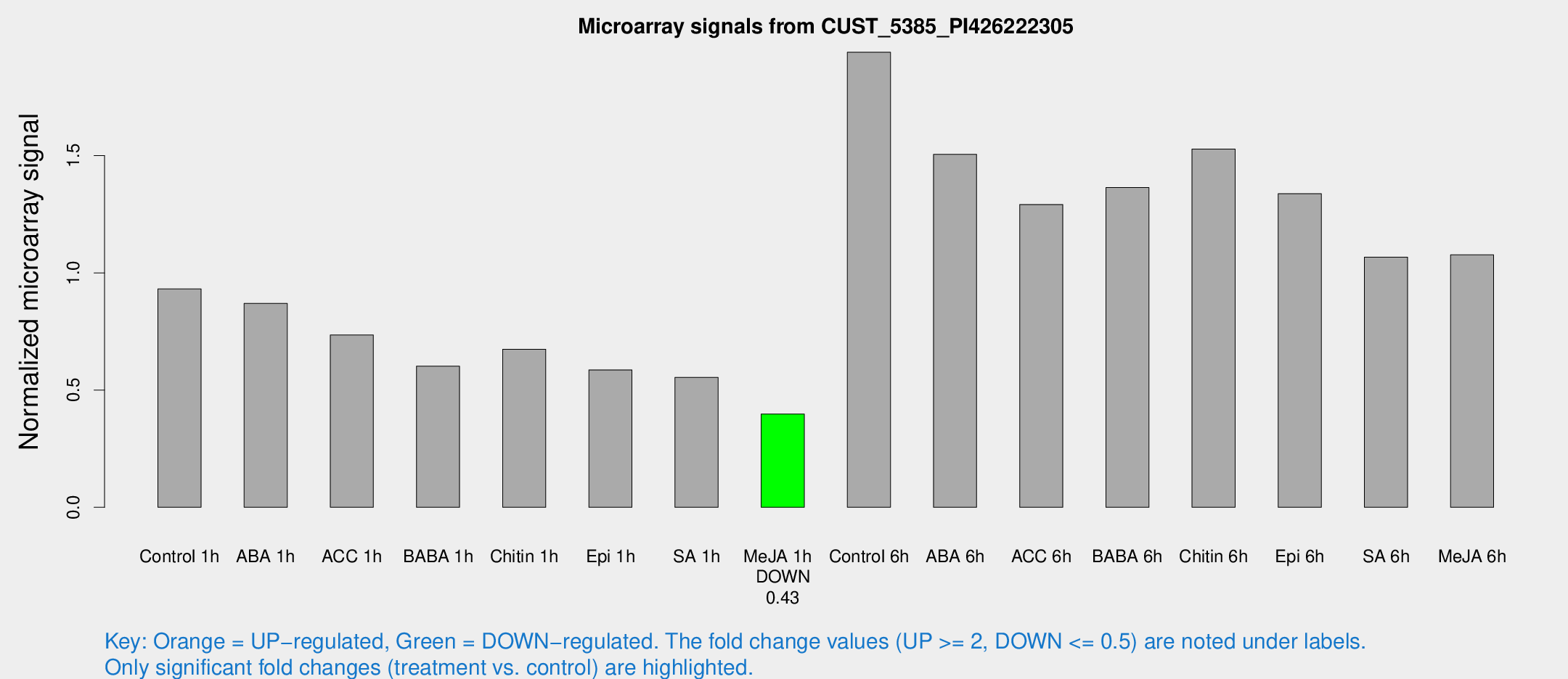
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 115.954 | 16.6773 | 0.931469 | 0.0742127 |
| ABA 1h | 94.4749 | 7.86309 | 0.869923 | 0.0579099 |
| ACC 1h | 98.8512 | 27.6735 | 0.735007 | 0.159652 |
| BABA 1h | 74.9884 | 16.3134 | 0.601686 | 0.0910535 |
| Chitin 1h | 74.6637 | 7.28096 | 0.674279 | 0.0484472 |
| Epi 1h | 62.4745 | 6.5368 | 0.586089 | 0.0452458 |
| SA 1h | 70.1626 | 7.52298 | 0.554202 | 0.0588944 |
| Me-JA 1h | 39.7436 | 4.00796 | 0.397773 | 0.0402218 |
| Control 6h | 260.235 | 72.4502 | 1.94098 | 0.477169 |
| ABA 6h | 195.542 | 11.8027 | 1.50522 | 0.0906864 |
| ACC 6h | 190.326 | 45.0342 | 1.29146 | 0.117001 |
| BABA 6h | 187.734 | 18.1583 | 1.36373 | 0.132857 |
| Chitin 6h | 201.847 | 26.2633 | 1.52776 | 0.163533 |
| Epi 6h | 186.586 | 23.6895 | 1.33727 | 0.179247 |
| SA 6h | 137.507 | 31.5097 | 1.06687 | 0.165472 |
| Me-JA 6h | 136.224 | 25.5796 | 1.07663 | 0.143164 |
Source Transcript PGSC0003DMT400003959 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G33255.1 | +3 | 1e-113 | 339 | 156/214 (73%) | Haloacid dehalogenase-like hydrolase (HAD) superfamily protein | chr2:14098795-14100358 FORWARD LENGTH=245 |
