Probe CUST_5363_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5363_PI426222305 | JHI_St_60k_v1 | DMT400009067 | GCTCGAGTGTTATTGGCTTCATTGATCCTTTTCTTAGTGCTATTCAATGTTATATGCAGC |
All Microarray Probes Designed to Gene DMG400003528
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5036_PI426222305 | JHI_St_60k_v1 | DMT400009066 | AGTGTTATTGGCTTCATTGATCCTTTTCTTAGTGCTATTCAATGTTATATGCAGCTAGTG |
| CUST_5363_PI426222305 | JHI_St_60k_v1 | DMT400009067 | GCTCGAGTGTTATTGGCTTCATTGATCCTTTTCTTAGTGCTATTCAATGTTATATGCAGC |
Microarray Signals from CUST_5363_PI426222305
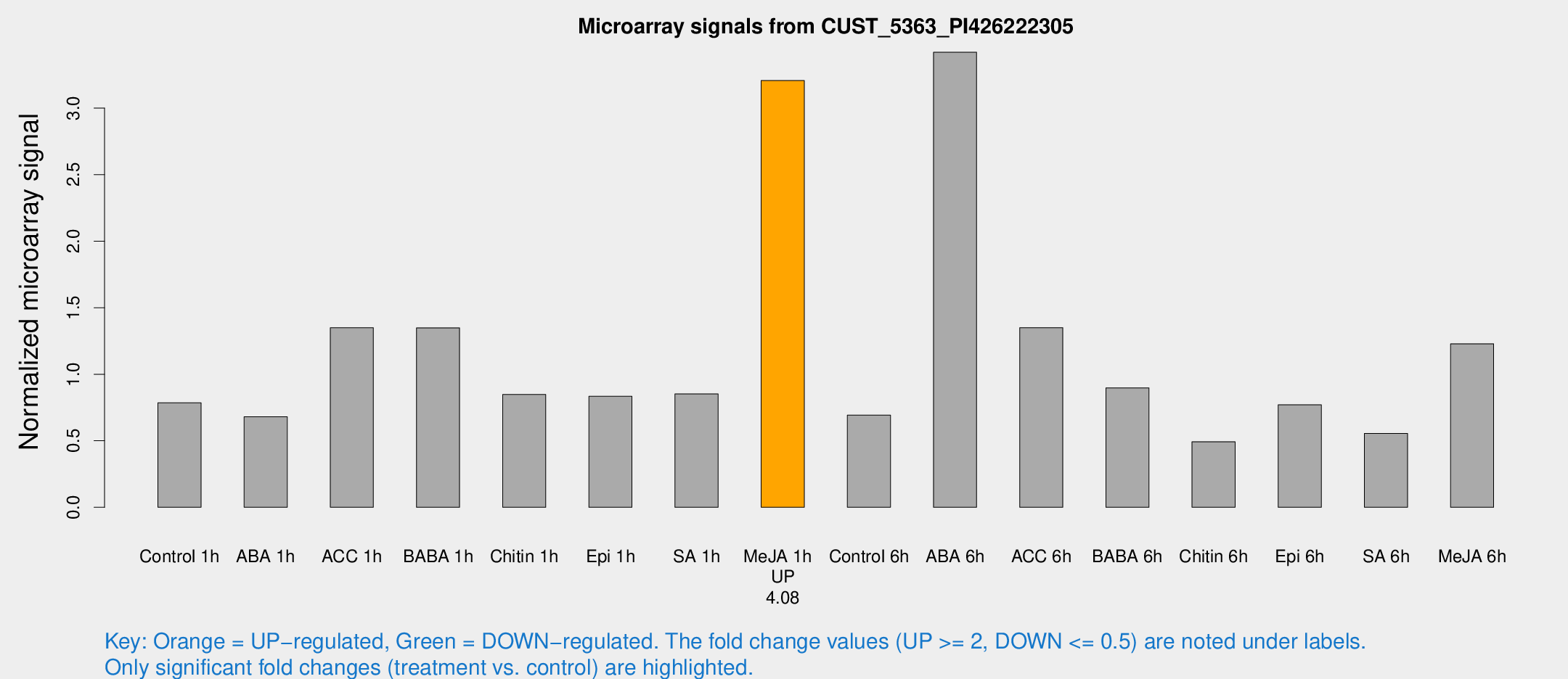
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25.2352 | 5.67108 | 0.785486 | 0.170036 |
| ABA 1h | 26.4132 | 15.108 | 0.680313 | 0.558252 |
| ACC 1h | 59.7953 | 26.3047 | 1.34949 | 1.13647 |
| BABA 1h | 39.7405 | 4.57522 | 1.34857 | 0.159432 |
| Chitin 1h | 24.3671 | 5.63645 | 0.847496 | 0.183683 |
| Epi 1h | 22.2424 | 3.87966 | 0.834154 | 0.153356 |
| SA 1h | 27.2637 | 4.96671 | 0.851583 | 0.140148 |
| Me-JA 1h | 79.6019 | 7.07087 | 3.20711 | 0.239798 |
| Control 6h | 24.2179 | 9.35855 | 0.693162 | 0.273037 |
| ABA 6h | 118.139 | 29.2169 | 3.41974 | 0.84124 |
| ACC 6h | 46.8454 | 5.35421 | 1.3502 | 0.15129 |
| BABA 6h | 30.6581 | 4.46482 | 0.897779 | 0.133201 |
| Chitin 6h | 18.1505 | 6.33045 | 0.492807 | 0.233965 |
| Epi 6h | 26.3425 | 4.95283 | 0.770365 | 0.148561 |
| SA 6h | 17.31 | 4.1902 | 0.555309 | 0.151349 |
| Me-JA 6h | 37.8019 | 5.5776 | 1.22866 | 0.145823 |
Source Transcript PGSC0003DMT400009067 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G18170.1 | +3 | 5e-126 | 381 | 176/226 (78%) | MAP kinase 7 | chr2:7908178-7909374 REVERSE LENGTH=368 |
