Probe CUST_5360_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5360_PI426222305 | JHI_St_60k_v1 | DMT400009126 | TTGATAGGACCTTTGAAGTCGCCAAGCAGTTTTTGTCTGGTGGACAATCTGAAGTTGAGG |
All Microarray Probes Designed to Gene DMG400003548
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5360_PI426222305 | JHI_St_60k_v1 | DMT400009126 | TTGATAGGACCTTTGAAGTCGCCAAGCAGTTTTTGTCTGGTGGACAATCTGAAGTTGAGG |
| CUST_5336_PI426222305 | JHI_St_60k_v1 | DMT400009127 | AATGCTTATAGCTGCTCCTCAAGGCTGTTTTGTTCATATACCGTGTGCTTTTAGCAATTT |
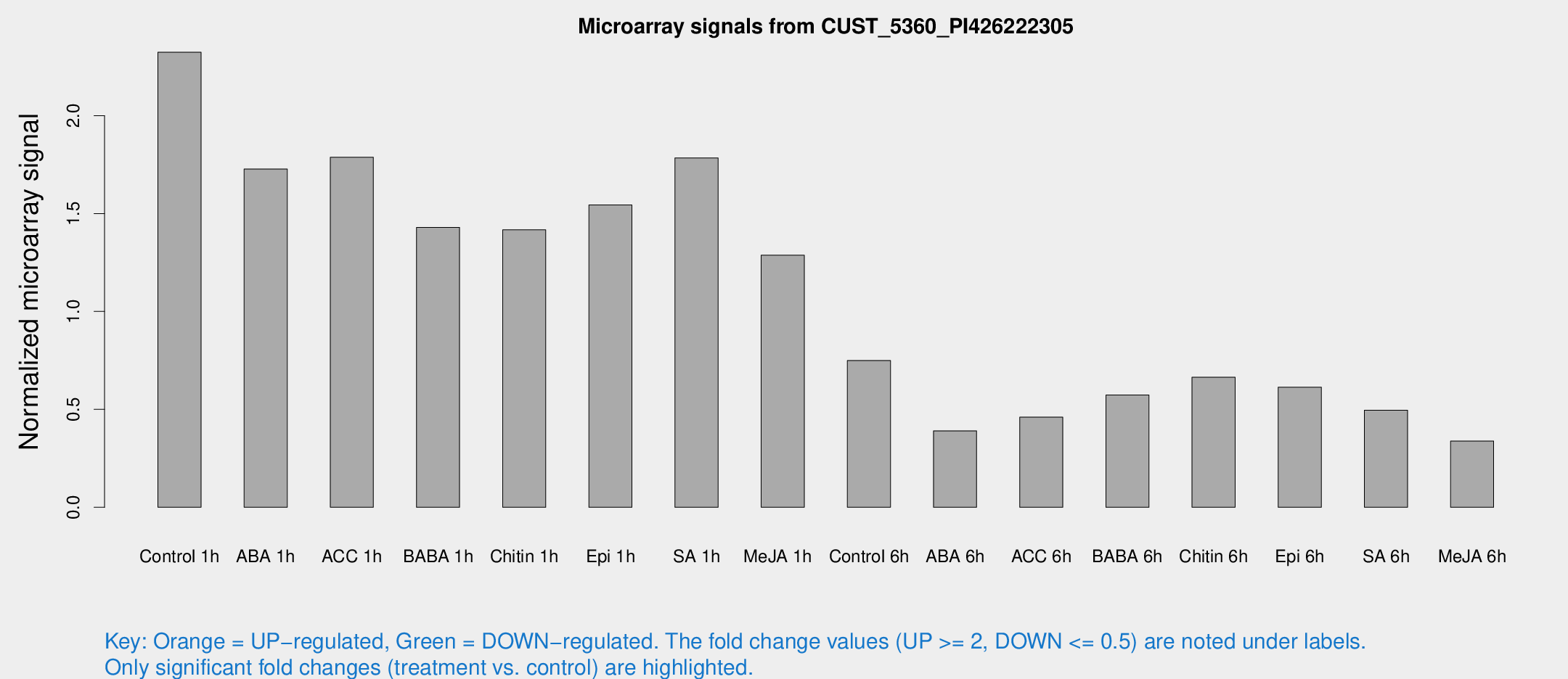
Microarray Signals from CUST_5360_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50676 | 10211.1 | 2.32371 | 0.304575 |
| ABA 1h | 32250.5 | 2235.72 | 1.72714 | 0.0997164 |
| ACC 1h | 39871.2 | 6783.46 | 1.78739 | 0.195877 |
| BABA 1h | 30425.4 | 6223.63 | 1.42929 | 0.1853 |
| Chitin 1h | 27275.4 | 3865.25 | 1.41682 | 0.095234 |
| Epi 1h | 28374 | 2815.52 | 1.54383 | 0.139058 |
| SA 1h | 39114.6 | 5071.44 | 1.78395 | 0.128727 |
| Me-JA 1h | 22111.8 | 1278.45 | 1.28729 | 0.0876484 |
| Control 6h | 16328.2 | 2801.4 | 0.749492 | 0.0768195 |
| ABA 6h | 8800.44 | 838.805 | 0.39044 | 0.0225426 |
| ACC 6h | 11436.8 | 1979.54 | 0.460156 | 0.033966 |
| BABA 6h | 13638.7 | 1494.18 | 0.573116 | 0.0713292 |
| Chitin 6h | 14971.5 | 1360.87 | 0.66418 | 0.0423973 |
| Epi 6h | 14539.2 | 840.039 | 0.613317 | 0.0519216 |
| SA 6h | 11038.4 | 2583.83 | 0.495496 | 0.080476 |
| Me-JA 6h | 7413.14 | 1525.49 | 0.338142 | 0.0466664 |
Source Transcript PGSC0003DMT400009126 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G38970.1 | +1 | 0.0 | 513 | 282/398 (71%) | fructose-bisphosphate aldolase 2 | chr4:18163714-18165659 REVERSE LENGTH=398 |
