Probe CUST_52832_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52832_PI426222305 | JHI_St_60k_v1 | DMT400083434 | AATTGTTTAGGACTGCAACAAAAATTCGGTCATAATGGACCTGATTGCGCAAGTCTTATA |
All Microarray Probes Designed to Gene DMG400033303
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52832_PI426222305 | JHI_St_60k_v1 | DMT400083434 | AATTGTTTAGGACTGCAACAAAAATTCGGTCATAATGGACCTGATTGCGCAAGTCTTATA |
Microarray Signals from CUST_52832_PI426222305
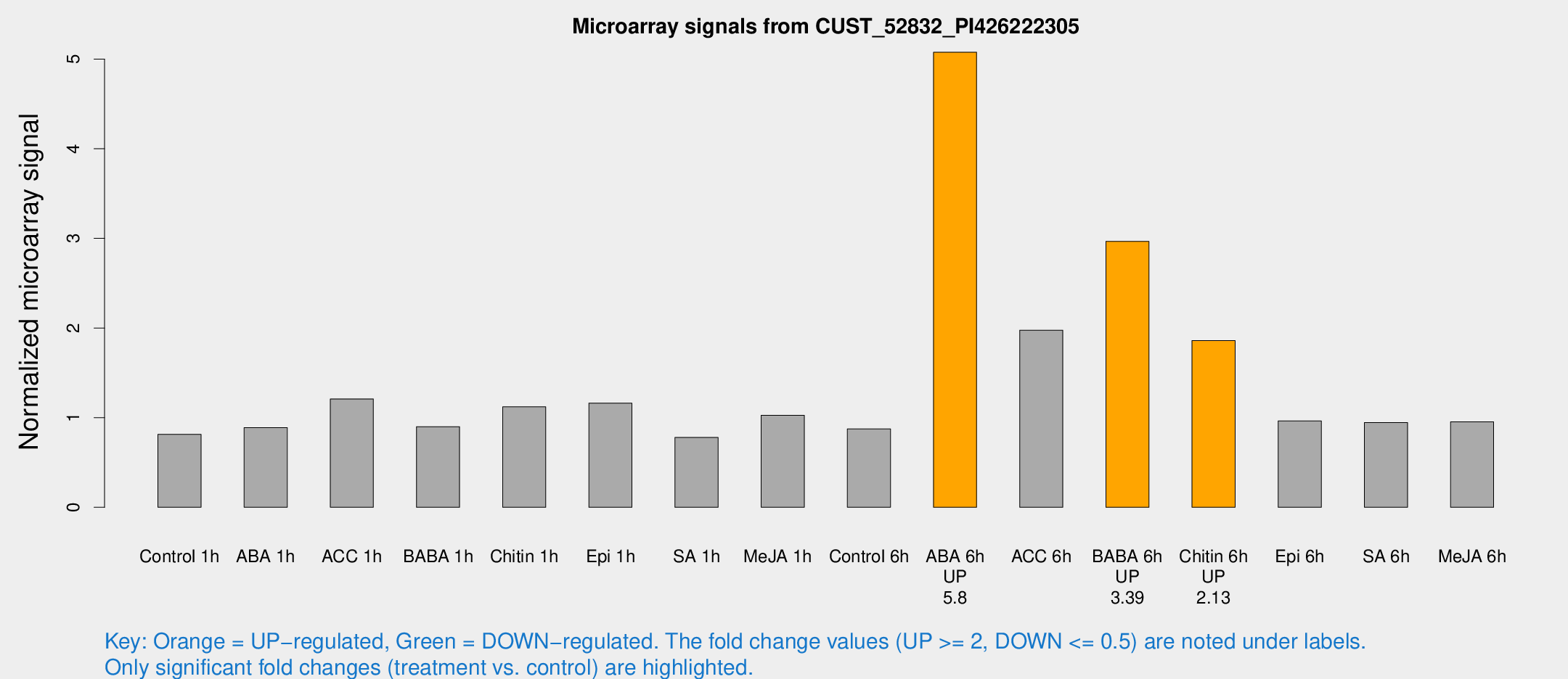
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.11432 | 3.54782 | 0.814782 | 0.472549 |
| ABA 1h | 5.89812 | 3.42374 | 0.888815 | 0.515903 |
| ACC 1h | 10.3944 | 3.90318 | 1.20908 | 0.568541 |
| BABA 1h | 6.51887 | 3.78933 | 0.89954 | 0.521731 |
| Chitin 1h | 7.92974 | 3.62528 | 1.12137 | 0.555468 |
| Epi 1h | 7.82266 | 3.4936 | 1.16241 | 0.568412 |
| SA 1h | 6.01589 | 3.49086 | 0.779244 | 0.451313 |
| Me-JA 1h | 6.3133 | 3.6817 | 1.02549 | 0.594503 |
| Control 6h | 6.58448 | 3.82019 | 0.874628 | 0.506797 |
| ABA 6h | 41.6621 | 7.10479 | 5.07661 | 0.811202 |
| ACC 6h | 20.7803 | 9.63975 | 1.97609 | 1.26797 |
| BABA 6h | 26.4381 | 6.784 | 2.9665 | 0.717441 |
| Chitin 6h | 15.284 | 4.44603 | 1.85972 | 0.573659 |
| Epi 6h | 8.21899 | 4.38839 | 0.963258 | 0.511084 |
| SA 6h | 7.02129 | 4.06896 | 0.946094 | 0.548135 |
| Me-JA 6h | 7.14155 | 3.87489 | 0.953653 | 0.519173 |
Source Transcript PGSC0003DMT400083434 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G23300.1 | +2 | 3e-09 | 54 | 24/44 (55%) | MATE efflux family protein | chr1:8263827-8266048 REVERSE LENGTH=515 |
