Probe CUST_52817_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52817_PI426222305 | JHI_St_60k_v1 | DMT400080992 | CACTACCTTTATGGCGCTTGGAACAACTTCTTTGATTGGTATGGAGGAATTCATGATAAC |
All Microarray Probes Designed to Gene DMG400031559
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52817_PI426222305 | JHI_St_60k_v1 | DMT400080992 | CACTACCTTTATGGCGCTTGGAACAACTTCTTTGATTGGTATGGAGGAATTCATGATAAC |
Microarray Signals from CUST_52817_PI426222305
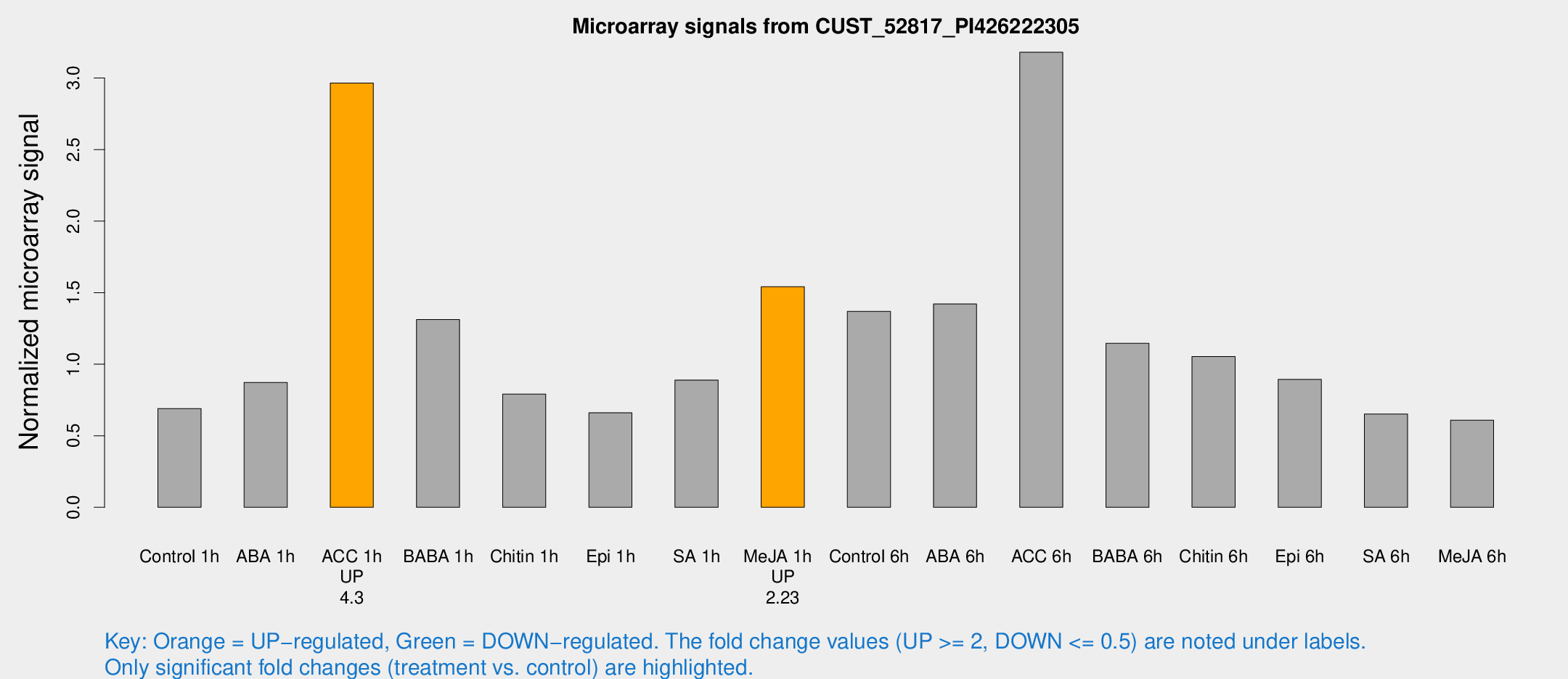
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.32617 | 3.48823 | 0.690035 | 0.351408 |
| ABA 1h | 8.13926 | 3.39722 | 0.872648 | 0.389236 |
| ACC 1h | 31.3198 | 4.78739 | 2.9643 | 0.481693 |
| BABA 1h | 12.9777 | 3.74693 | 1.31185 | 0.383911 |
| Chitin 1h | 7.41447 | 3.46716 | 0.791049 | 0.393281 |
| Epi 1h | 5.83005 | 3.38417 | 0.660479 | 0.382626 |
| SA 1h | 11.2369 | 5.11796 | 0.889197 | 0.427805 |
| Me-JA 1h | 13.4006 | 3.69146 | 1.54194 | 0.472205 |
| Control 6h | 17.8269 | 8.63063 | 1.36936 | 0.687514 |
| ABA 6h | 15.6341 | 3.77482 | 1.42061 | 0.356175 |
| ACC 6h | 38.3033 | 6.31029 | 3.18027 | 0.532377 |
| BABA 6h | 15.1333 | 5.40794 | 1.14573 | 0.471798 |
| Chitin 6h | 13.9615 | 6.53836 | 1.05299 | 0.527566 |
| Epi 6h | 10.8373 | 4.33668 | 0.893728 | 0.399571 |
| SA 6h | 6.565 | 3.8048 | 0.651548 | 0.377302 |
| Me-JA 6h | 6.17966 | 3.49975 | 0.6093 | 0.345314 |
Source Transcript PGSC0003DMT400080992 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14520.1 | +3 | 2e-06 | 36 | 17/54 (31%) | Terpenoid cyclases/Protein prenyltransferases superfamily protein | chr3:4875868-4878243 REVERSE LENGTH=605 |
