Probe CUST_52661_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52661_PI426222305 | JHI_St_60k_v1 | DMT400020094 | GAACATATCAGGTAATTAACTGACTCCCATCTTTCTTTTCAATTGTGCTGCCACTGGTAA |
All Microarray Probes Designed to Gene DMG400007782
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52661_PI426222305 | JHI_St_60k_v1 | DMT400020094 | GAACATATCAGGTAATTAACTGACTCCCATCTTTCTTTTCAATTGTGCTGCCACTGGTAA |
Microarray Signals from CUST_52661_PI426222305
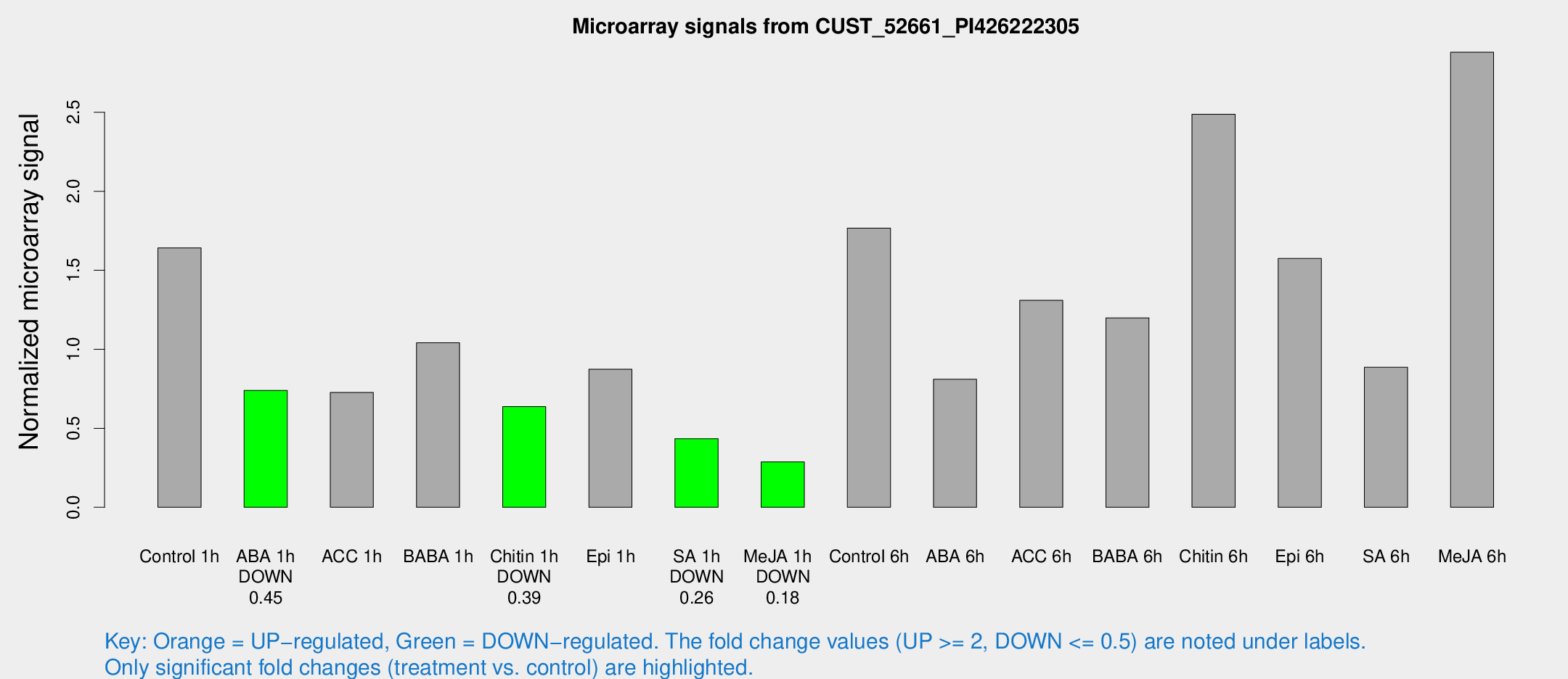
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 60.601 | 9.06268 | 1.64182 | 0.139131 |
| ABA 1h | 24.1572 | 3.75194 | 0.740064 | 0.136584 |
| ACC 1h | 31.3118 | 10.334 | 0.72665 | 0.26869 |
| BABA 1h | 38.2572 | 9.33756 | 1.04138 | 0.203109 |
| Chitin 1h | 21.8879 | 4.69542 | 0.637395 | 0.127073 |
| Epi 1h | 27.4412 | 3.90355 | 0.873522 | 0.12397 |
| SA 1h | 16.332 | 3.81251 | 0.434215 | 0.10479 |
| Me-JA 1h | 8.93112 | 3.79705 | 0.287414 | 0.138092 |
| Control 6h | 72.437 | 26.3695 | 1.76718 | 0.563286 |
| ABA 6h | 36.6595 | 12.6219 | 0.810442 | 0.368768 |
| ACC 6h | 54.7523 | 5.49819 | 1.30962 | 0.129474 |
| BABA 6h | 55.4382 | 21.3588 | 1.19906 | 0.452776 |
| Chitin 6h | 98.3736 | 15.5161 | 2.48722 | 0.44407 |
| Epi 6h | 67.9718 | 17.2214 | 1.57468 | 0.408522 |
| SA 6h | 32.0531 | 4.40284 | 0.886371 | 0.153361 |
| Me-JA 6h | 110.58 | 29.6593 | 2.88003 | 0.585998 |
Source Transcript PGSC0003DMT400020094 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G29320.1 | +2 | 0.0 | 704 | 350/471 (74%) | Glycosyl transferase, family 35 | chr3:11252871-11257587 FORWARD LENGTH=962 |
