Probe CUST_52549_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52549_PI426222305 | JHI_St_60k_v1 | DMT400082143 | CATCAAGCGAATTGTATTTAGGATTCTCTTTCGGCAATTTAAAGCCATACACACCTTGTA |
All Microarray Probes Designed to Gene DMG400032292
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52549_PI426222305 | JHI_St_60k_v1 | DMT400082143 | CATCAAGCGAATTGTATTTAGGATTCTCTTTCGGCAATTTAAAGCCATACACACCTTGTA |
Microarray Signals from CUST_52549_PI426222305
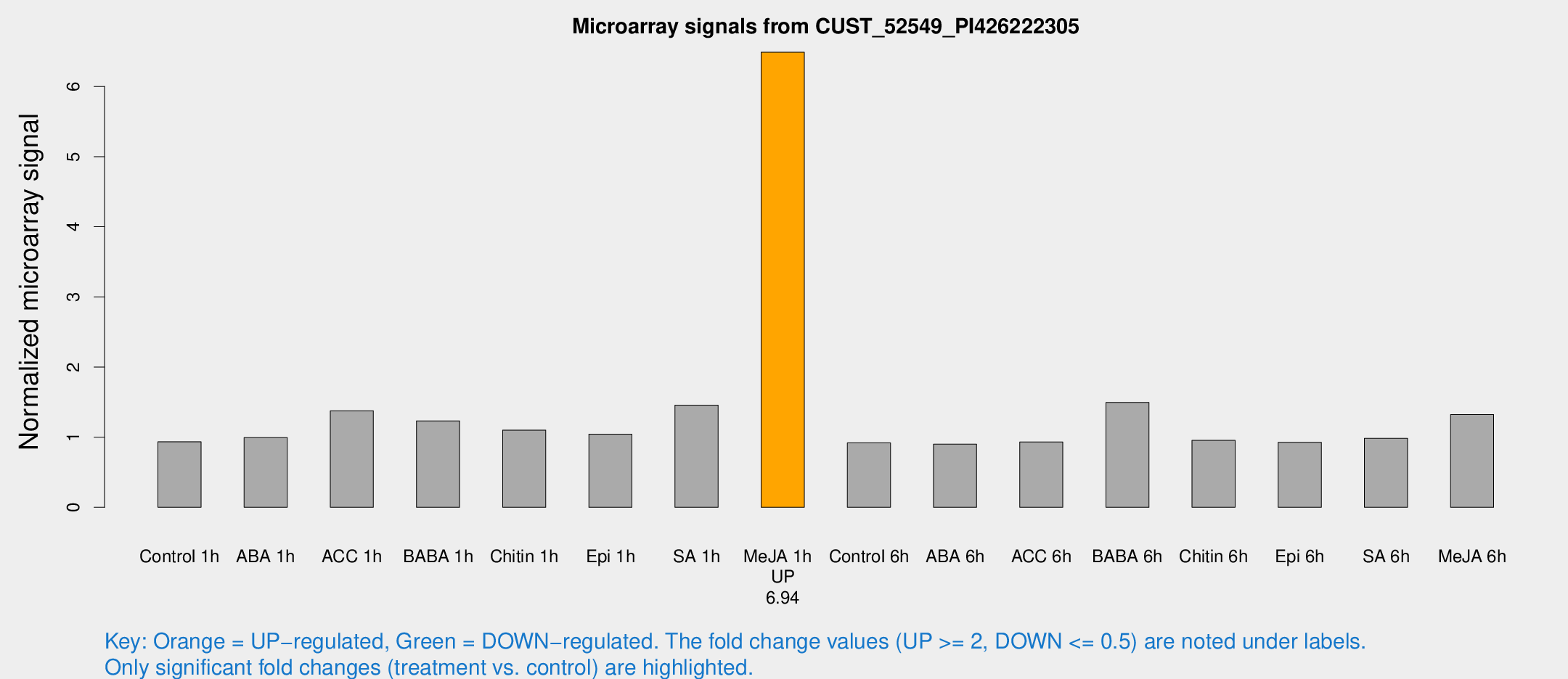
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.34725 | 2.9159 | 0.934386 | 0.512028 |
| ABA 1h | 4.86087 | 2.81764 | 0.993656 | 0.557078 |
| ACC 1h | 8.39738 | 3.19708 | 1.37693 | 0.576741 |
| BABA 1h | 6.91218 | 3.11396 | 1.23245 | 0.574603 |
| Chitin 1h | 5.55244 | 2.9598 | 1.10079 | 0.574182 |
| Epi 1h | 5.06402 | 2.93973 | 1.04338 | 0.592844 |
| SA 1h | 9.29762 | 2.9885 | 1.45664 | 0.567814 |
| Me-JA 1h | 30.9788 | 4.94116 | 6.48813 | 0.752539 |
| Control 6h | 5.25717 | 3.04565 | 0.918181 | 0.531857 |
| ABA 6h | 5.47071 | 3.17355 | 0.899849 | 0.521395 |
| ACC 6h | 6.24578 | 3.70513 | 0.931889 | 0.539887 |
| BABA 6h | 11.2781 | 4.76543 | 1.49448 | 0.623453 |
| Chitin 6h | 5.8031 | 3.36286 | 0.954264 | 0.552897 |
| Epi 6h | 6.00943 | 3.50993 | 0.926049 | 0.536587 |
| SA 6h | 5.55603 | 3.21956 | 0.982716 | 0.569072 |
| Me-JA 6h | 7.74244 | 3.08069 | 1.32257 | 0.563072 |
Source Transcript PGSC0003DMT400082143 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G20340.1 | +1 | 1e-07 | 49 | 29/61 (48%) | Expression of the gene is downregulated in the presence of paraquat, an inducer of photoxidative stress. | chr3:7093075-7093422 REVERSE LENGTH=115 |
