Probe CUST_52443_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52443_PI426222305 | JHI_St_60k_v1 | DMT400049146 | GTTAGCATCAGCATGACACCTGATTATTTCGATTATTGTGCAGCACTAGGGTGGCAAATG |
All Microarray Probes Designed to Gene DMG400019100
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52442_PI426222305 | JHI_St_60k_v1 | DMT400049147 | CATTGGTGGGATTGCTTGCTATTGGAGTTAATTTTCGGTACTCGAGTTTTGAGAATTGTC |
| CUST_52443_PI426222305 | JHI_St_60k_v1 | DMT400049146 | GTTAGCATCAGCATGACACCTGATTATTTCGATTATTGTGCAGCACTAGGGTGGCAAATG |
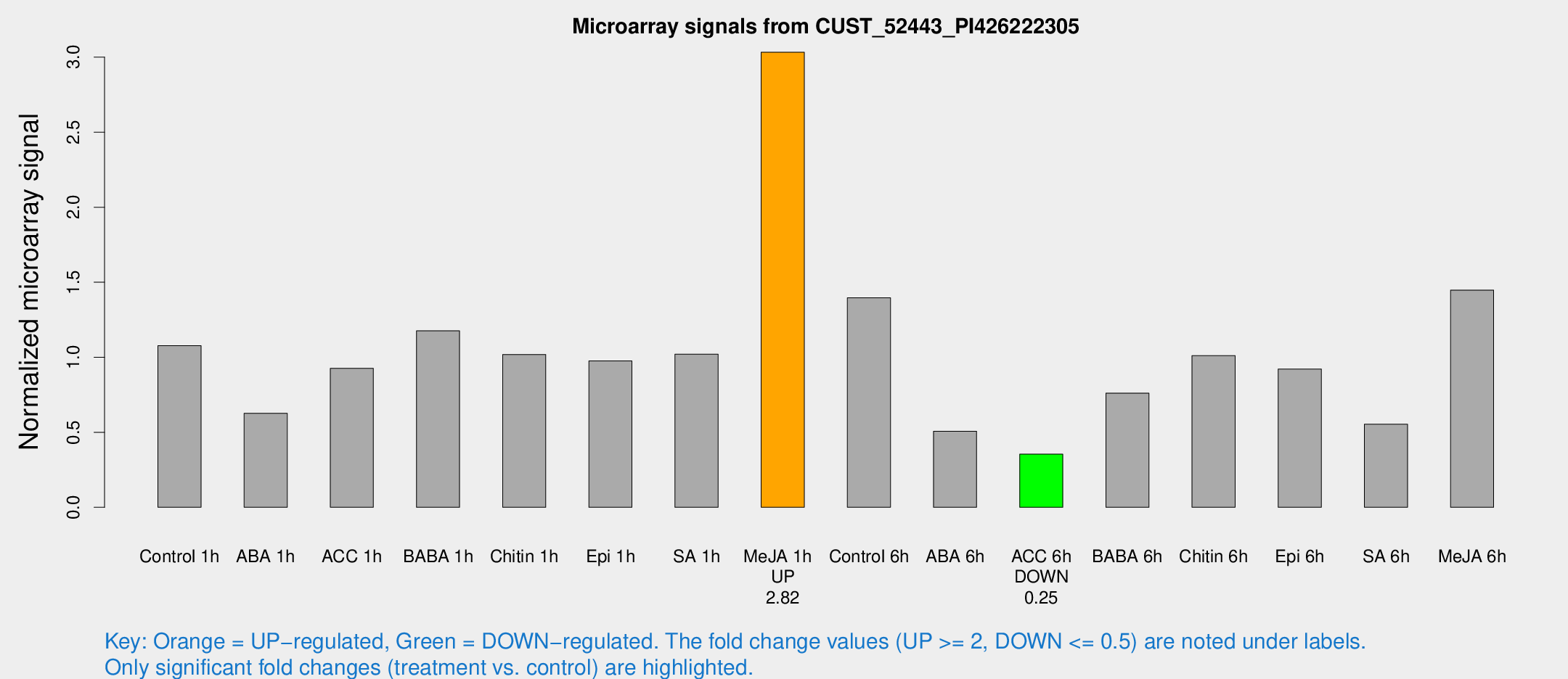
Microarray Signals from CUST_52443_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 346.527 | 79.9587 | 1.07703 | 0.170387 |
| ABA 1h | 178.573 | 39.0807 | 0.626731 | 0.0992776 |
| ACC 1h | 308.02 | 65.6228 | 0.926322 | 0.160099 |
| BABA 1h | 360.813 | 66.274 | 1.17612 | 0.121336 |
| Chitin 1h | 282.417 | 22.4072 | 1.01859 | 0.128136 |
| Epi 1h | 261.297 | 24.9829 | 0.975496 | 0.080655 |
| SA 1h | 338.287 | 72.8882 | 1.02066 | 0.297784 |
| Me-JA 1h | 787.111 | 152.868 | 3.03281 | 0.318229 |
| Control 6h | 476.98 | 139.037 | 1.39631 | 0.363096 |
| ABA 6h | 165.582 | 10.0603 | 0.506846 | 0.0307256 |
| ACC 6h | 127.424 | 20.1572 | 0.354109 | 0.0226316 |
| BABA 6h | 289.977 | 96.6347 | 0.760791 | 0.245505 |
| Chitin 6h | 330.954 | 20.2628 | 1.0105 | 0.0925568 |
| Epi 6h | 360.676 | 121.998 | 0.92081 | 0.419892 |
| SA 6h | 206.849 | 98.3476 | 0.553549 | 0.234626 |
| Me-JA 6h | 491.631 | 137.95 | 1.44761 | 0.429364 |
Source Transcript PGSC0003DMT400049146 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G23560.1 | +3 | 2e-109 | 268 | 139/225 (62%) | MATE efflux family protein | chr3:8454361-8456588 REVERSE LENGTH=477 |
