Probe CUST_52442_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52442_PI426222305 | JHI_St_60k_v1 | DMT400049147 | CATTGGTGGGATTGCTTGCTATTGGAGTTAATTTTCGGTACTCGAGTTTTGAGAATTGTC |
All Microarray Probes Designed to Gene DMG400019100
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52442_PI426222305 | JHI_St_60k_v1 | DMT400049147 | CATTGGTGGGATTGCTTGCTATTGGAGTTAATTTTCGGTACTCGAGTTTTGAGAATTGTC |
| CUST_52443_PI426222305 | JHI_St_60k_v1 | DMT400049146 | GTTAGCATCAGCATGACACCTGATTATTTCGATTATTGTGCAGCACTAGGGTGGCAAATG |
Microarray Signals from CUST_52442_PI426222305
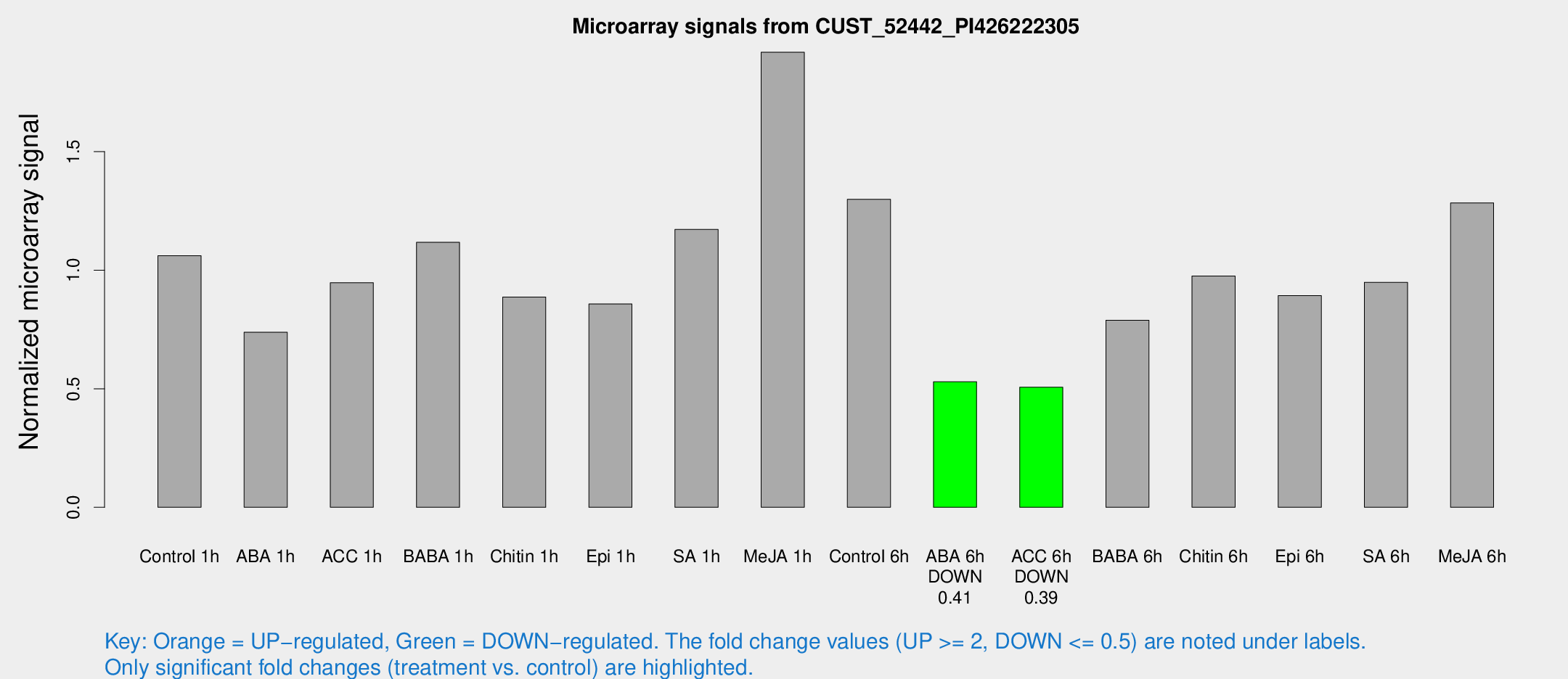
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 17068 | 2564.4 | 1.0614 | 0.0975143 |
| ABA 1h | 10773.2 | 2441.55 | 0.738019 | 0.11167 |
| ACC 1h | 16490.8 | 4138.46 | 0.94731 | 0.207255 |
| BABA 1h | 17137.9 | 1724.28 | 1.11757 | 0.0645237 |
| Chitin 1h | 12688 | 1286.61 | 0.887007 | 0.088788 |
| Epi 1h | 11776.6 | 1023.25 | 0.857349 | 0.0579661 |
| SA 1h | 21646.3 | 7828.12 | 1.17212 | 0.519819 |
| Me-JA 1h | 24863.1 | 2164.29 | 1.91942 | 0.110818 |
| Control 6h | 21998 | 5828.52 | 1.29916 | 0.243147 |
| ABA 6h | 8966.17 | 938.288 | 0.529642 | 0.0744324 |
| ACC 6h | 9408.85 | 1646.12 | 0.506279 | 0.029231 |
| BABA 6h | 14230.2 | 2213.51 | 0.789004 | 0.108989 |
| Chitin 6h | 16463.5 | 1320.67 | 0.975771 | 0.100954 |
| Epi 6h | 17122.3 | 4177.95 | 0.893209 | 0.356931 |
| SA 6h | 14890.6 | 1226.88 | 0.949091 | 0.0547964 |
| Me-JA 6h | 20751.3 | 3414.66 | 1.28391 | 0.122836 |
Source Transcript PGSC0003DMT400049147 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G23560.1 | +1 | 5e-170 | 495 | 272/444 (61%) | MATE efflux family protein | chr3:8454361-8456588 REVERSE LENGTH=477 |
