Probe CUST_52315_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52315_PI426222305 | JHI_St_60k_v1 | DMT400041422 | ACATACATATATCGATAGGCTTGTGGAATATATCGGTGTGGGAGTGGAAGCAGCGAAGGA |
All Microarray Probes Designed to Gene DMG400016051
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_52315_PI426222305 | JHI_St_60k_v1 | DMT400041422 | ACATACATATATCGATAGGCTTGTGGAATATATCGGTGTGGGAGTGGAAGCAGCGAAGGA |
Microarray Signals from CUST_52315_PI426222305
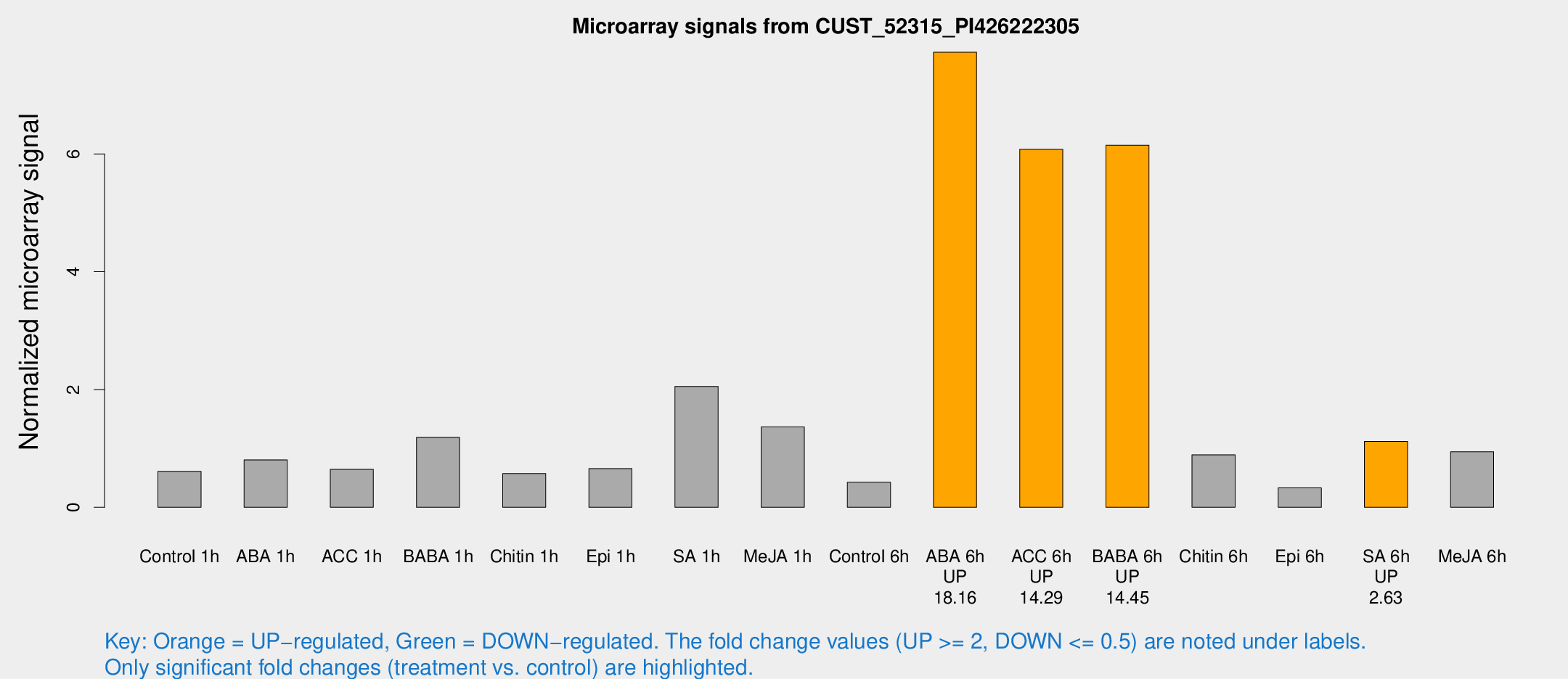
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 74.3814 | 11.675 | 0.609672 | 0.0993642 |
| ABA 1h | 87.4812 | 16.5515 | 0.805563 | 0.153272 |
| ACC 1h | 83.3536 | 18.5305 | 0.644004 | 0.118814 |
| BABA 1h | 142.287 | 27.693 | 1.18746 | 0.140172 |
| Chitin 1h | 69.6175 | 21.9547 | 0.572752 | 0.236616 |
| Epi 1h | 73.8078 | 20.9026 | 0.657994 | 0.208842 |
| SA 1h | 263.687 | 54.9756 | 2.05271 | 0.373406 |
| Me-JA 1h | 142.627 | 40.0512 | 1.3639 | 0.260598 |
| Control 6h | 55.3588 | 14.5903 | 0.425588 | 0.0999637 |
| ABA 6h | 1082.49 | 300.923 | 7.72724 | 2.44559 |
| ACC 6h | 837.903 | 91.2576 | 6.08083 | 1.30821 |
| BABA 6h | 884.284 | 249.029 | 6.14905 | 1.72119 |
| Chitin 6h | 131.713 | 49.6941 | 0.892213 | 0.387414 |
| Epi 6h | 58.4584 | 28.8112 | 0.329411 | 0.186846 |
| SA 6h | 141.041 | 33.8871 | 1.11806 | 0.187783 |
| Me-JA 6h | 116.911 | 25.4054 | 0.94311 | 0.187121 |
Source Transcript PGSC0003DMT400041422 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49690.1 | +1 | 4e-55 | 191 | 140/469 (30%) | UDP-Glycosyltransferase superfamily protein | chr5:20189968-20191350 REVERSE LENGTH=460 |
