Probe CUST_51754_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51754_PI426222305 | JHI_St_60k_v1 | DMT400041450 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
All Microarray Probes Designed to Gene DMG400016067
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51753_PI426222305 | JHI_St_60k_v1 | DMT400041452 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
| CUST_51754_PI426222305 | JHI_St_60k_v1 | DMT400041450 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
Microarray Signals from CUST_51754_PI426222305
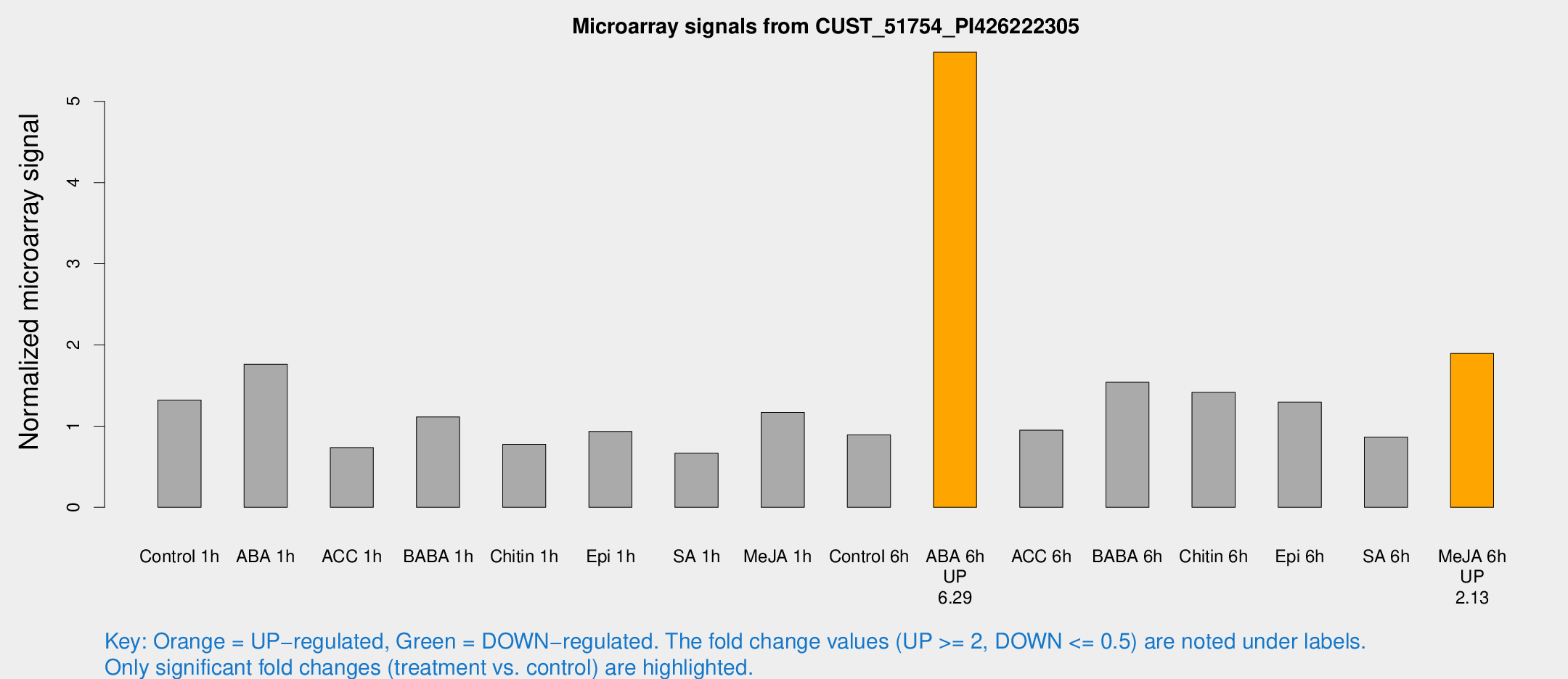
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11.3655 | 3.83246 | 1.31992 | 0.589973 |
| ABA 1h | 13.8829 | 4.52029 | 1.76214 | 0.753126 |
| ACC 1h | 5.77326 | 3.35122 | 0.73599 | 0.426219 |
| BABA 1h | 8.6197 | 3.19005 | 1.11339 | 0.461641 |
| Chitin 1h | 5.19264 | 3.02379 | 0.775382 | 0.437553 |
| Epi 1h | 6.23731 | 2.9862 | 0.934428 | 0.462619 |
| SA 1h | 5.22572 | 3.00427 | 0.667121 | 0.383138 |
| Me-JA 1h | 7.92299 | 3.11727 | 1.16863 | 0.551002 |
| Control 6h | 6.94114 | 3.1576 | 0.891487 | 0.424298 |
| ABA 6h | 46.9641 | 8.8917 | 5.60561 | 0.801925 |
| ACC 6h | 8.30161 | 3.83924 | 0.949342 | 0.430967 |
| BABA 6h | 14.2913 | 4.23774 | 1.54048 | 0.482016 |
| Chitin 6h | 11.7173 | 3.51418 | 1.41619 | 0.443021 |
| Epi 6h | 12.3059 | 3.68623 | 1.29723 | 0.477898 |
| SA 6h | 6.633 | 3.29114 | 0.865404 | 0.445796 |
| Me-JA 6h | 15.4839 | 4.29614 | 1.89541 | 0.462234 |
Source Transcript PGSC0003DMT400041450 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14710.5 | +3 | 3e-19 | 69 | 41/95 (43%) | RmlC-like cupins superfamily protein | chr4:8424897-8426174 REVERSE LENGTH=201 |
