Probe CUST_51753_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51753_PI426222305 | JHI_St_60k_v1 | DMT400041452 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
All Microarray Probes Designed to Gene DMG400016067
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51753_PI426222305 | JHI_St_60k_v1 | DMT400041452 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
| CUST_51754_PI426222305 | JHI_St_60k_v1 | DMT400041450 | CAACACGATGTTTTTGTTATCTGATTTGGCCATCACTTGCATGCATATTAGCAATTTTTC |
Microarray Signals from CUST_51753_PI426222305
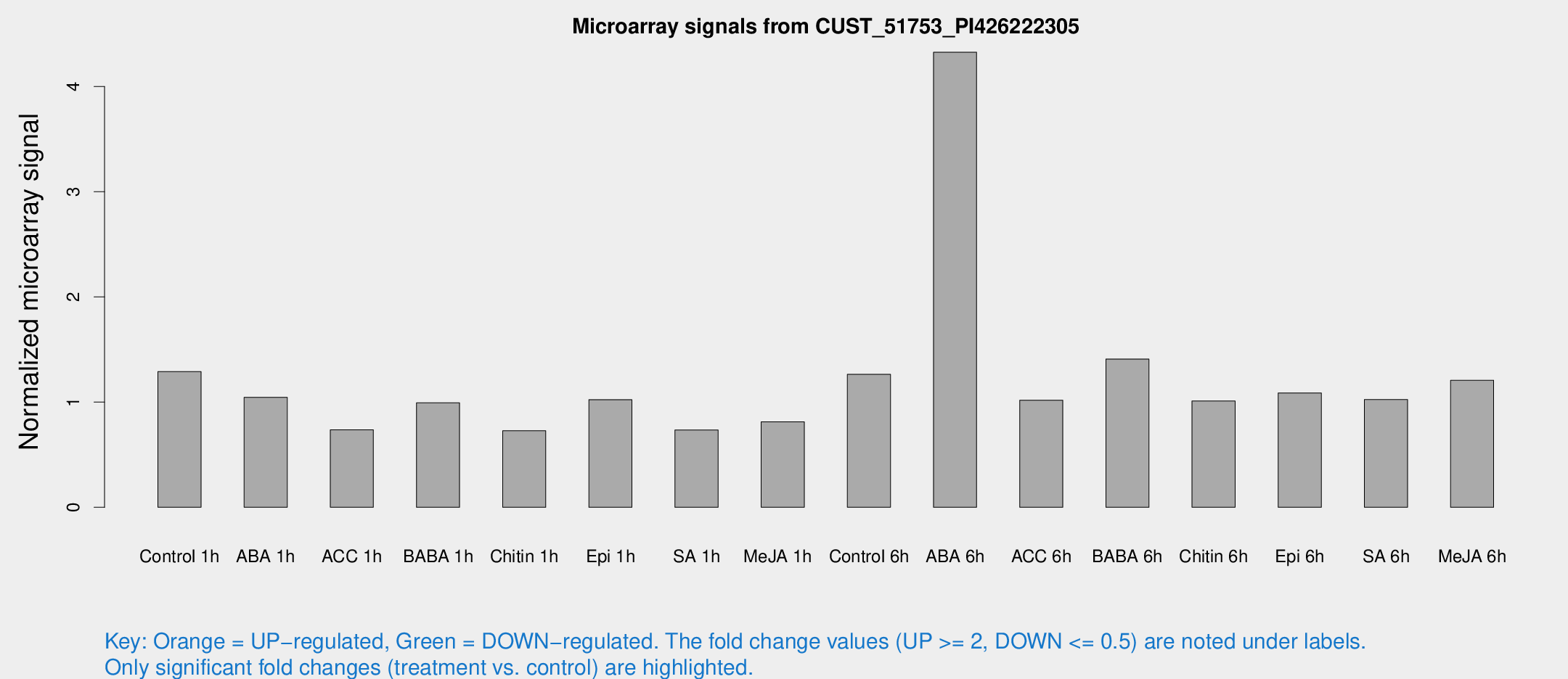
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10.8368 | 3.18239 | 1.29068 | 0.389003 |
| ABA 1h | 7.94559 | 3.03831 | 1.04591 | 0.433074 |
| ACC 1h | 6.30533 | 3.63013 | 0.73705 | 0.420557 |
| BABA 1h | 8.20796 | 3.29852 | 0.993349 | 0.427816 |
| Chitin 1h | 5.36036 | 3.11533 | 0.727368 | 0.416086 |
| Epi 1h | 7.4748 | 3.09417 | 1.02339 | 0.448994 |
| SA 1h | 6.44917 | 3.13274 | 0.734856 | 0.377082 |
| Me-JA 1h | 5.5081 | 3.20237 | 0.813108 | 0.471193 |
| Control 6h | 11.4046 | 3.27708 | 1.2639 | 0.456408 |
| ABA 6h | 38.3424 | 3.99294 | 4.32656 | 0.59243 |
| ACC 6h | 10.9859 | 4.01177 | 1.01864 | 0.484732 |
| BABA 6h | 13.1602 | 3.64531 | 1.40963 | 0.40383 |
| Chitin 6h | 9.15335 | 3.5885 | 1.01019 | 0.427124 |
| Epi 6h | 11.9 | 4.96319 | 1.08687 | 0.450148 |
| SA 6h | 8.62095 | 3.40256 | 1.02437 | 0.426489 |
| Me-JA 6h | 11.4591 | 4.19859 | 1.20729 | 0.462003 |
Source Transcript PGSC0003DMT400041452 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14710.5 | +1 | 1e-28 | 70 | 31/40 (78%) | RmlC-like cupins superfamily protein | chr4:8424897-8426174 REVERSE LENGTH=201 |
