Probe CUST_51624_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51624_PI426222305 | JHI_St_60k_v1 | DMT400053854 | TTTAGGACAGGGGAATTACTGTCTTGAAAACTCTAGCAGAGGTGTACATAACAGAAAAAA |
All Microarray Probes Designed to Gene DMG400020896
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51624_PI426222305 | JHI_St_60k_v1 | DMT400053854 | TTTAGGACAGGGGAATTACTGTCTTGAAAACTCTAGCAGAGGTGTACATAACAGAAAAAA |
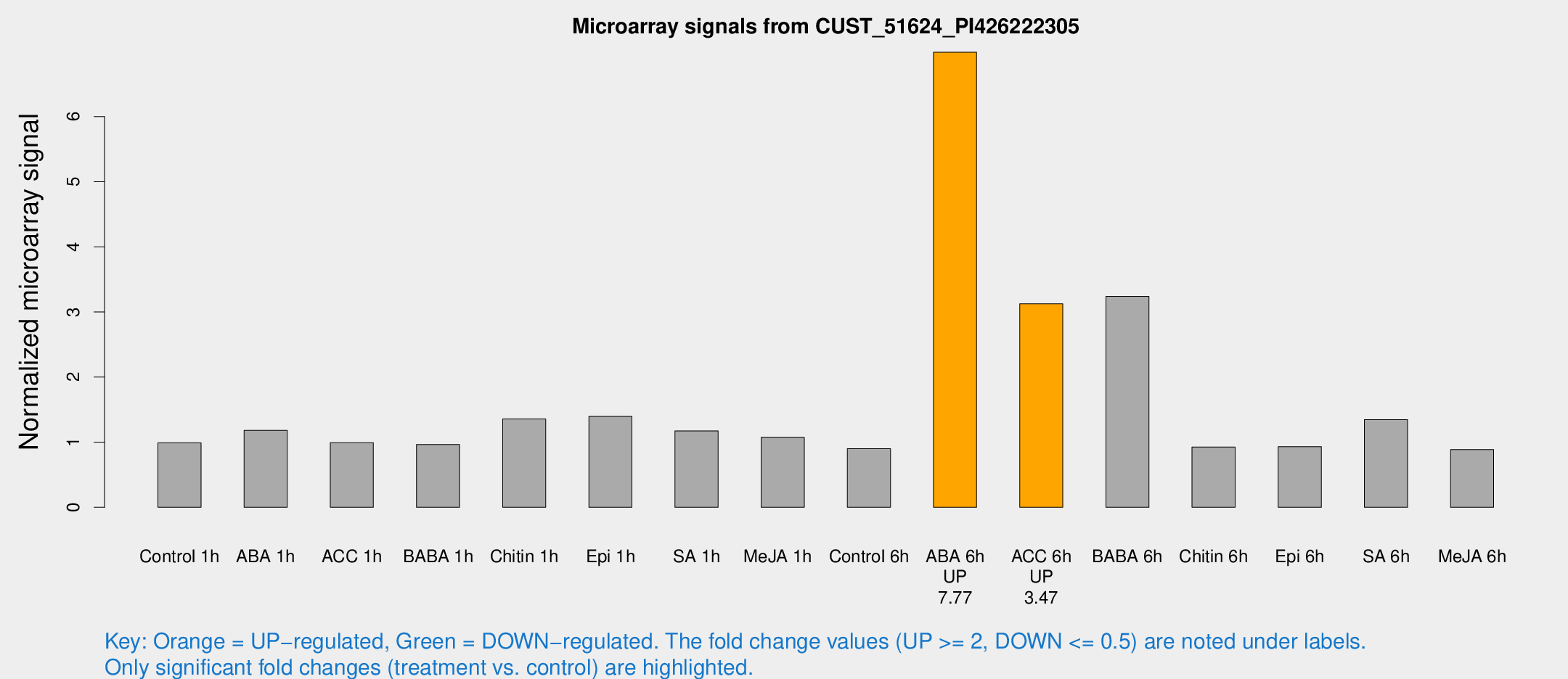
Microarray Signals from CUST_51624_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.40385 | 3.21147 | 0.988902 | 0.515361 |
| ABA 1h | 6.67992 | 3.06046 | 1.18253 | 0.558589 |
| ACC 1h | 6.47073 | 3.73543 | 0.992084 | 0.567116 |
| BABA 1h | 5.87117 | 3.40765 | 0.963721 | 0.558094 |
| Chitin 1h | 8.25516 | 3.21782 | 1.35691 | 0.627182 |
| Epi 1h | 8.73147 | 3.51334 | 1.39595 | 0.646271 |
| SA 1h | 8.72594 | 3.41095 | 1.17332 | 0.553265 |
| Me-JA 1h | 5.56219 | 3.24225 | 1.07381 | 0.62202 |
| Control 6h | 5.69168 | 3.30223 | 0.899387 | 0.521195 |
| ABA 6h | 54.2683 | 17.6985 | 6.98874 | 2.80242 |
| ACC 6h | 22.9038 | 4.14021 | 3.12451 | 0.553266 |
| BABA 6h | 32.4443 | 19.4037 | 3.24032 | 2.25166 |
| Chitin 6h | 6.22559 | 3.60908 | 0.926577 | 0.536702 |
| Epi 6h | 6.69771 | 3.78516 | 0.931876 | 0.520674 |
| SA 6h | 9.50212 | 3.52519 | 1.34546 | 0.731791 |
| Me-JA 6h | 5.57239 | 3.23159 | 0.887146 | 0.514398 |
Source Transcript PGSC0003DMT400053854 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G49690.1 | +3 | 1e-63 | 220 | 154/482 (32%) | UDP-Glycosyltransferase superfamily protein | chr5:20189968-20191350 REVERSE LENGTH=460 |
