Probe CUST_51616_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51616_PI426222305 | JHI_St_60k_v1 | DMT400053851 | TTGTGACTCTCACGGCTGCTCGTGGAACTATTGAATATGTTGCTCCAGAGTTGATTAGCA |
All Microarray Probes Designed to Gene DMG400020894
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51619_PI426222305 | JHI_St_60k_v1 | DMT400053850 | AGTTGGCAGAGGAAGTATGACATTATTCTTGGAGTGGCTCGGAGGAATTGGGTATTTGCA |
| CUST_51616_PI426222305 | JHI_St_60k_v1 | DMT400053851 | TTGTGACTCTCACGGCTGCTCGTGGAACTATTGAATATGTTGCTCCAGAGTTGATTAGCA |
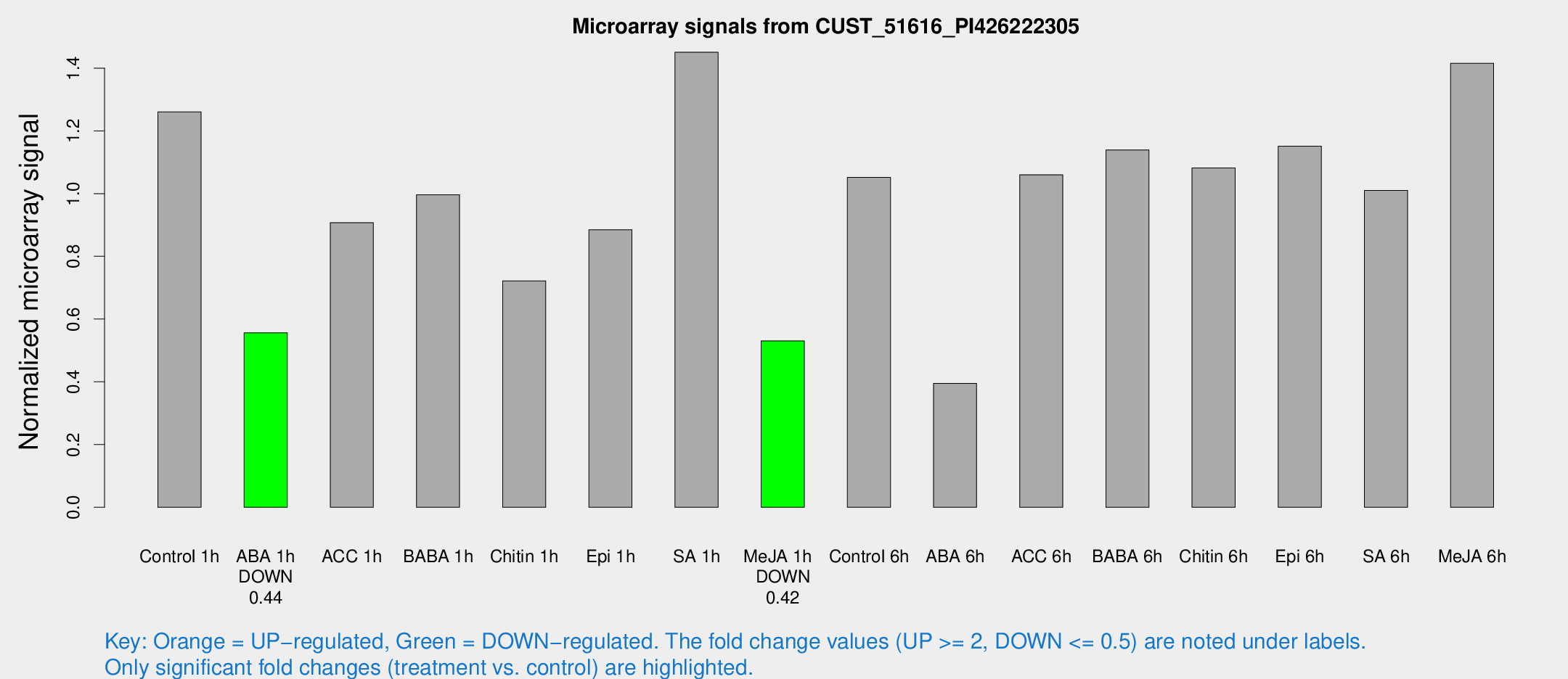
Microarray Signals from CUST_51616_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 11107.7 | 1893.95 | 1.26034 | 0.131789 |
| ABA 1h | 4297.26 | 565.065 | 0.556294 | 0.0359066 |
| ACC 1h | 9303.25 | 2957.24 | 0.907442 | 0.345038 |
| BABA 1h | 8644.05 | 1760.38 | 0.996097 | 0.126146 |
| Chitin 1h | 5621.58 | 561.877 | 0.721784 | 0.0416744 |
| Epi 1h | 6596.98 | 410.041 | 0.884851 | 0.0525557 |
| SA 1h | 12952.4 | 1457.43 | 1.45066 | 0.0837548 |
| Me-JA 1h | 3825.09 | 670.706 | 0.530563 | 0.0465239 |
| Control 6h | 9696.6 | 2348.09 | 1.05161 | 0.206071 |
| ABA 6h | 3729.68 | 646.714 | 0.394901 | 0.0579857 |
| ACC 6h | 10512.6 | 931.625 | 1.06012 | 0.0612073 |
| BABA 6h | 11182 | 1556.94 | 1.13916 | 0.129727 |
| Chitin 6h | 9970.96 | 952.329 | 1.08155 | 0.096243 |
| Epi 6h | 11458.7 | 1960.26 | 1.15093 | 0.170038 |
| SA 6h | 9344.47 | 2687.16 | 1.01004 | 0.207603 |
| Me-JA 6h | 12461.7 | 2110.02 | 1.41512 | 0.12881 |
Source Transcript PGSC0003DMT400053851 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G38280.1 | +3 | 9e-59 | 204 | 100/209 (48%) | PR5-like receptor kinase | chr5:15293325-15295838 REVERSE LENGTH=665 |
