Probe CUST_51335_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51335_PI426222305 | JHI_St_60k_v1 | DMT400028134 | TGTTCTTCATATGTTCACTTTTGCTTGTACGTTGTATATAGGTGGTATTTCACATATGGG |
All Microarray Probes Designed to Gene DMG400010855
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51336_PI426222305 | JHI_St_60k_v1 | DMT400028137 | AGAAGGAAGCACAGACGATTCTAAATAGACCTCTGTCACAGGTGGTATTTCACATATGGG |
| CUST_51335_PI426222305 | JHI_St_60k_v1 | DMT400028134 | TGTTCTTCATATGTTCACTTTTGCTTGTACGTTGTATATAGGTGGTATTTCACATATGGG |
Microarray Signals from CUST_51335_PI426222305
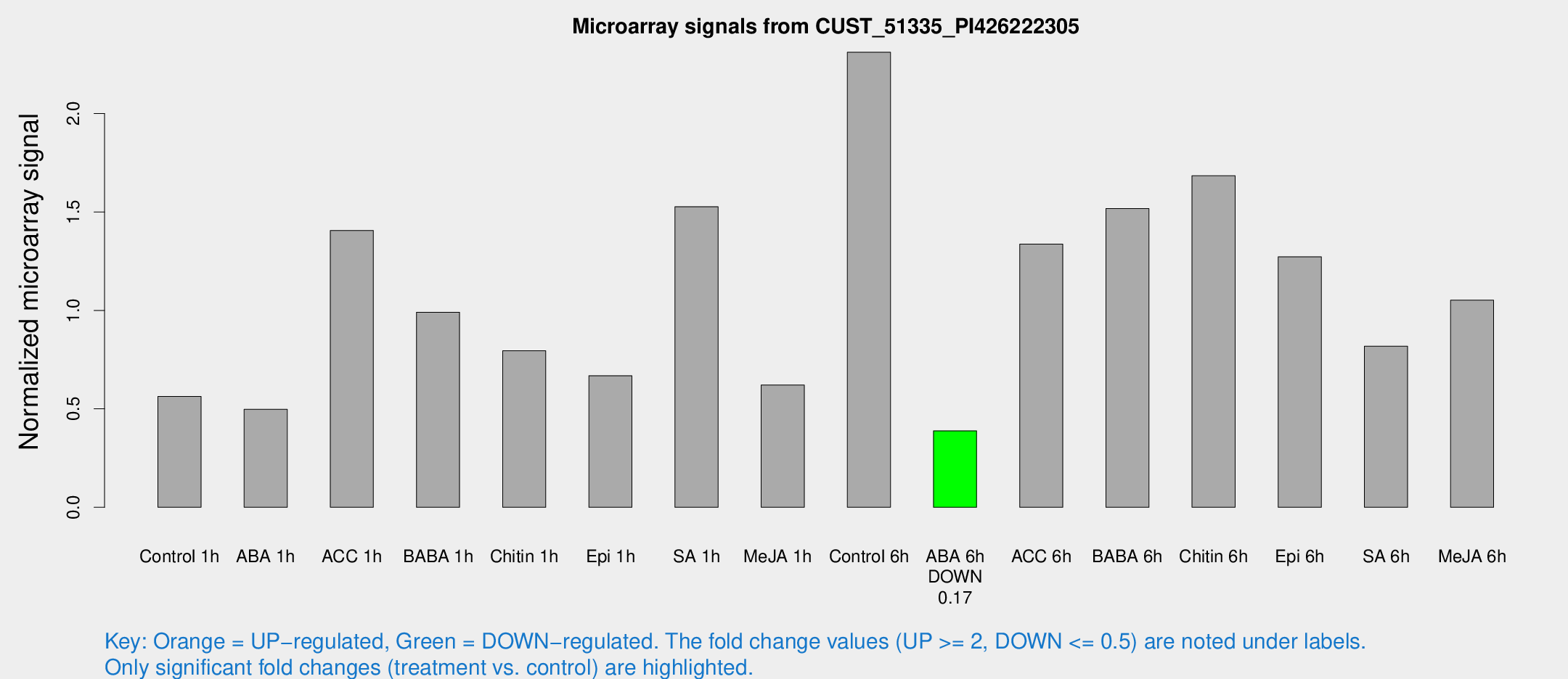
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.10007 | 3.78771 | 0.563096 | 0.239858 |
| ABA 1h | 7.21391 | 3.63876 | 0.497757 | 0.259198 |
| ACC 1h | 25.8534 | 7.71436 | 1.406 | 0.447473 |
| BABA 1h | 18.4968 | 7.29906 | 0.990625 | 0.692405 |
| Chitin 1h | 15.5978 | 8.88376 | 0.795194 | 0.540783 |
| Epi 1h | 11.2841 | 5.23988 | 0.668035 | 0.346162 |
| SA 1h | 25.7353 | 4.03308 | 1.52646 | 0.343511 |
| Me-JA 1h | 8.15562 | 3.83557 | 0.621265 | 0.292426 |
| Control 6h | 42.1205 | 12.5388 | 2.31103 | 0.720634 |
| ABA 6h | 6.6307 | 3.85102 | 0.388415 | 0.225047 |
| ACC 6h | 26.8437 | 7.76783 | 1.33652 | 0.299755 |
| BABA 6h | 27.4487 | 4.44676 | 1.51718 | 0.24699 |
| Chitin 6h | 30.8623 | 7.33922 | 1.68366 | 0.460046 |
| Epi 6h | 25.5301 | 7.52232 | 1.27204 | 0.601967 |
| SA 6h | 13.7879 | 4.10403 | 0.818466 | 0.289417 |
| Me-JA 6h | 20.6521 | 7.31764 | 1.05163 | 0.529815 |
Source Transcript PGSC0003DMT400028134 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52200.2 | +1 | 7e-12 | 65 | 34/52 (65%) | phosphoprotein phosphatase inhibitors | chr5:21202818-21204017 REVERSE LENGTH=211 |
