Probe CUST_51148_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51148_PI426222305 | JHI_St_60k_v1 | DMT400008556 | CTATCACAGTAAAACACTATTAGATCTGCTGATGTTGCAACTTGATTTTCGAAACTAAGC |
All Microarray Probes Designed to Gene DMG400003308
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51148_PI426222305 | JHI_St_60k_v1 | DMT400008556 | CTATCACAGTAAAACACTATTAGATCTGCTGATGTTGCAACTTGATTTTCGAAACTAAGC |
| CUST_51155_PI426222305 | JHI_St_60k_v1 | DMT400008555 | CTATCACAGTAAAACACTATTAGATCTGCTGATGTTGCAACTTGATTTTCGAAACTAAGC |
Microarray Signals from CUST_51148_PI426222305
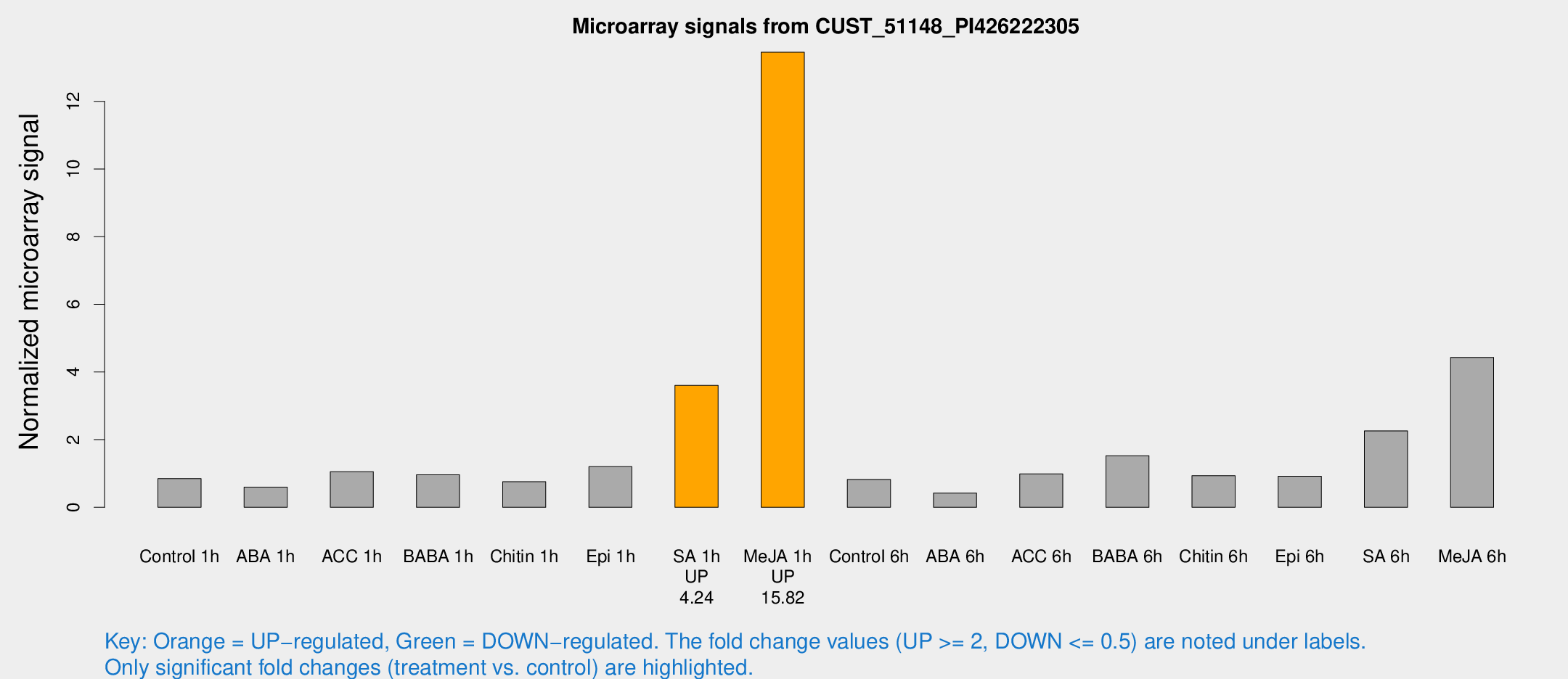
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.6839 | 3.37607 | 0.850445 | 0.155177 |
| ABA 1h | 13.2782 | 4.15981 | 0.597569 | 0.2629 |
| ACC 1h | 28.2042 | 9.48107 | 1.05317 | 0.448334 |
| BABA 1h | 23.2239 | 8.83128 | 0.963648 | 0.520286 |
| Chitin 1h | 15.1803 | 3.43869 | 0.756905 | 0.17773 |
| Epi 1h | 23.7474 | 4.59086 | 1.20109 | 0.292735 |
| SA 1h | 90.7883 | 30.3285 | 3.60339 | 1.46462 |
| Me-JA 1h | 240.585 | 14.2981 | 13.45 | 1.24978 |
| Control 6h | 25.5899 | 13.9016 | 0.82397 | 0.538584 |
| ABA 6h | 10.3341 | 3.621 | 0.421479 | 0.167044 |
| ACC 6h | 33.658 | 18.4365 | 0.988106 | 0.813575 |
| BABA 6h | 41.1027 | 11.9774 | 1.5262 | 0.442024 |
| Chitin 6h | 24.3983 | 8.06454 | 0.933429 | 0.333117 |
| Epi 6h | 25.7268 | 9.23883 | 0.918494 | 0.386712 |
| SA 6h | 51.2882 | 10.0541 | 2.25951 | 0.75414 |
| Me-JA 6h | 97.6587 | 9.63232 | 4.42993 | 0.602163 |
Source Transcript PGSC0003DMT400008556 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G37320.1 | +1 | 3e-62 | 207 | 107/202 (53%) | cytochrome P450, family 81, subfamily D, polypeptide 5 | chr4:17559742-17561690 REVERSE LENGTH=495 |
