Probe CUST_51005_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51005_PI426222305 | JHI_St_60k_v1 | DMT400034108 | TTTACCCTCCCGATAGTTACCCCGAGTCCTAATTGTTGTATTTTCGAAGTTGGTTTGAAG |
All Microarray Probes Designed to Gene DMG400013103
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_51005_PI426222305 | JHI_St_60k_v1 | DMT400034108 | TTTACCCTCCCGATAGTTACCCCGAGTCCTAATTGTTGTATTTTCGAAGTTGGTTTGAAG |
Microarray Signals from CUST_51005_PI426222305
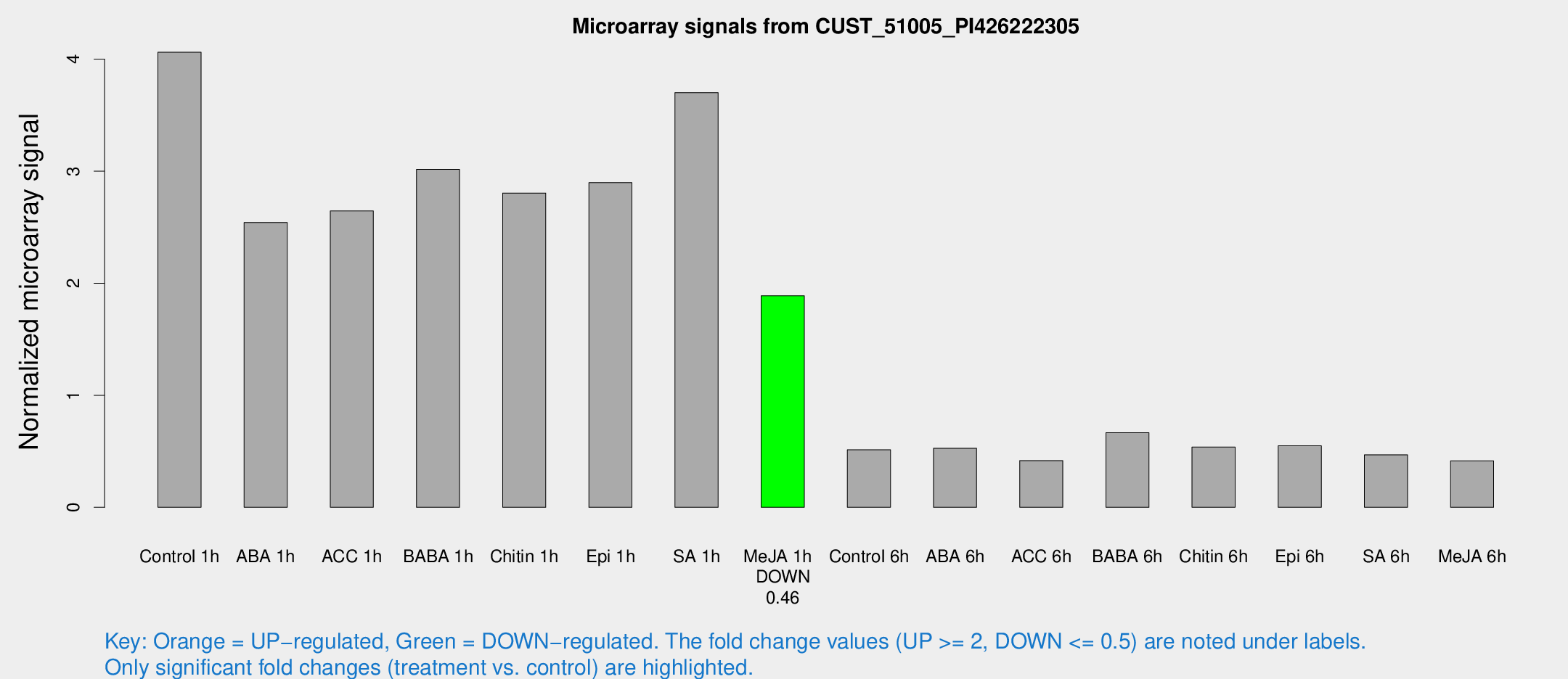
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6875.33 | 498.489 | 4.06176 | 0.234516 |
| ABA 1h | 3838.29 | 451.027 | 2.5424 | 0.146805 |
| ACC 1h | 5355.2 | 1745.38 | 2.64524 | 1.03182 |
| BABA 1h | 5087.78 | 944.316 | 3.01599 | 0.312393 |
| Chitin 1h | 4258.63 | 246.565 | 2.80479 | 0.255266 |
| Epi 1h | 4225.01 | 244.038 | 2.89759 | 0.16731 |
| SA 1h | 6424.56 | 376.803 | 3.70101 | 0.251463 |
| Me-JA 1h | 2619.52 | 252.176 | 1.88856 | 0.109069 |
| Control 6h | 884.041 | 119.836 | 0.513663 | 0.0379692 |
| ABA 6h | 984.119 | 183.743 | 0.527326 | 0.0889102 |
| ACC 6h | 807.285 | 46.9206 | 0.417138 | 0.0594134 |
| BABA 6h | 1259.91 | 76.3685 | 0.665826 | 0.0385092 |
| Chitin 6h | 968.572 | 56.263 | 0.538522 | 0.0378856 |
| Epi 6h | 1066.28 | 150.835 | 0.549768 | 0.0594033 |
| SA 6h | 830.485 | 207.505 | 0.468551 | 0.0773387 |
| Me-JA 6h | 744.201 | 192.215 | 0.414425 | 0.0757993 |
Source Transcript PGSC0003DMT400034108 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G20693.1 | +1 | 1e-28 | 108 | 51/102 (50%) | high mobility group B2 | chr1:7177282-7178487 FORWARD LENGTH=144 |
