Probe CUST_50798_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50798_PI426222305 | JHI_St_60k_v1 | DMT400027833 | GGAAGAAAGCATGGGGAATTTAGAAAAGATTGCACTATGCATCATATGTGATTTTTGTGT |
All Microarray Probes Designed to Gene DMG400010716
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50798_PI426222305 | JHI_St_60k_v1 | DMT400027833 | GGAAGAAAGCATGGGGAATTTAGAAAAGATTGCACTATGCATCATATGTGATTTTTGTGT |
Microarray Signals from CUST_50798_PI426222305
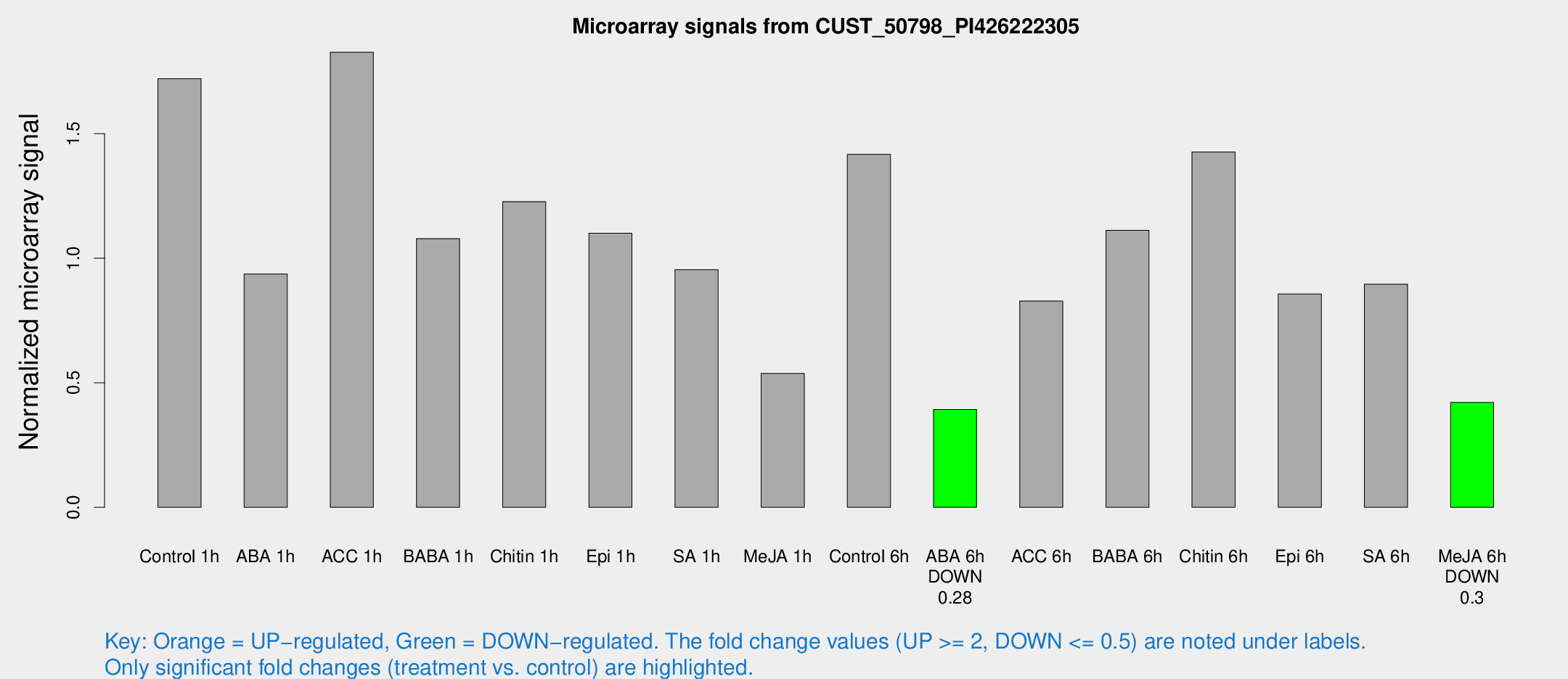
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 231.073 | 57.6552 | 1.72086 | 0.419484 |
| ABA 1h | 108.479 | 22.5601 | 0.936692 | 0.235858 |
| ACC 1h | 244.765 | 49.2803 | 1.82709 | 0.519254 |
| BABA 1h | 134.719 | 25.2924 | 1.07894 | 0.116866 |
| Chitin 1h | 143.602 | 31.3872 | 1.22718 | 0.249923 |
| Epi 1h | 120.783 | 17.1127 | 1.10033 | 0.168081 |
| SA 1h | 124.429 | 18.3074 | 0.953865 | 0.0842961 |
| Me-JA 1h | 54.8057 | 4.77133 | 0.537394 | 0.0885424 |
| Control 6h | 194.586 | 56.9362 | 1.41744 | 0.364397 |
| ABA 6h | 52.4183 | 5.1675 | 0.393035 | 0.0370599 |
| ACC 6h | 126.139 | 32.9778 | 0.828018 | 0.108135 |
| BABA 6h | 175.916 | 66.6051 | 1.11193 | 0.415227 |
| Chitin 6h | 191.546 | 24.0299 | 1.4267 | 0.132176 |
| Epi 6h | 126.978 | 27.6462 | 0.856612 | 0.296692 |
| SA 6h | 111.524 | 13.3962 | 0.895628 | 0.0990821 |
| Me-JA 6h | 57.448 | 16.7026 | 0.421077 | 0.105484 |
Source Transcript PGSC0003DMT400027833 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G34060.1 | +2 | 1e-37 | 140 | 65/103 (63%) | Peroxidase superfamily protein | chr2:14384914-14386530 FORWARD LENGTH=346 |
