Probe CUST_50508_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50508_PI426222305 | JHI_St_60k_v1 | DMT400062092 | TTAGTGAGAAGTTCTCTAAGCGCCTAGCCTGTGAGCAACACAATAGCGAAGTCTTTGTTA |
All Microarray Probes Designed to Gene DMG402024163
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50499_PI426222305 | JHI_St_60k_v1 | DMT400062093 | GCACATCACATCCGATGAATATCCTTTGTTTATTAGTCCACTTGTTCCATATTATCTGTC |
| CUST_50508_PI426222305 | JHI_St_60k_v1 | DMT400062092 | TTAGTGAGAAGTTCTCTAAGCGCCTAGCCTGTGAGCAACACAATAGCGAAGTCTTTGTTA |
Microarray Signals from CUST_50508_PI426222305
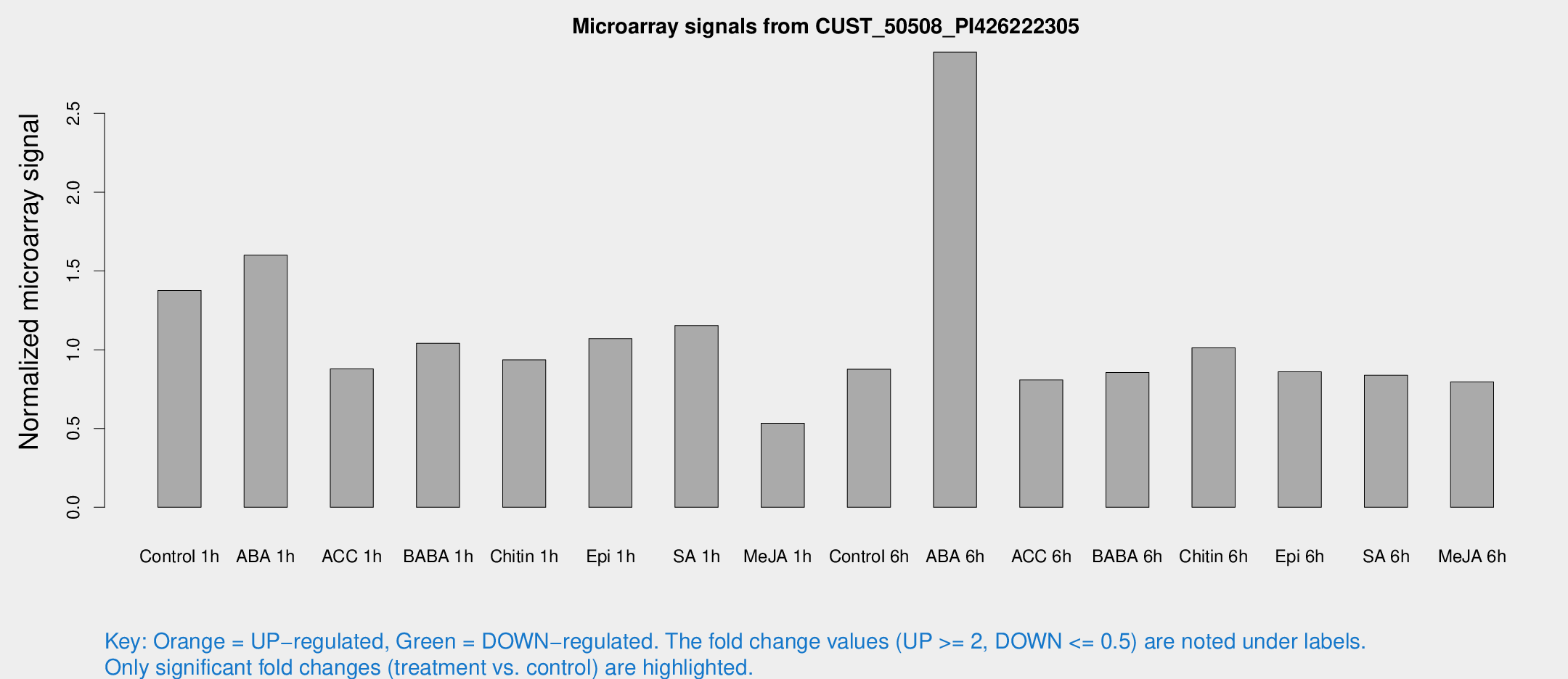
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 271.066 | 35.6349 | 1.37639 | 0.120235 |
| ABA 1h | 276.664 | 29.1557 | 1.59973 | 0.094414 |
| ACC 1h | 190.28 | 49.8504 | 0.87815 | 0.214921 |
| BABA 1h | 235.563 | 90.263 | 1.04087 | 0.432803 |
| Chitin 1h | 165.933 | 24.4202 | 0.936119 | 0.0805955 |
| Epi 1h | 179.949 | 13.0222 | 1.07072 | 0.0954609 |
| SA 1h | 240.356 | 50.7618 | 1.15368 | 0.19202 |
| Me-JA 1h | 86.7155 | 15.5441 | 0.533336 | 0.0683833 |
| Control 6h | 178.244 | 40.2271 | 0.876301 | 0.152122 |
| ABA 6h | 631.812 | 141.264 | 2.88829 | 0.657101 |
| ACC 6h | 180.362 | 15.2177 | 0.808295 | 0.0502832 |
| BABA 6h | 192.056 | 37.9644 | 0.855931 | 0.172586 |
| Chitin 6h | 208.379 | 12.6572 | 1.01269 | 0.061493 |
| Epi 6h | 188.44 | 13.9649 | 0.86046 | 0.0708055 |
| SA 6h | 174.347 | 50.7888 | 0.837856 | 0.174018 |
| Me-JA 6h | 161.096 | 36.279 | 0.796174 | 0.113971 |
Source Transcript PGSC0003DMT400062092 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17550.1 | +1 | 1e-74 | 237 | 119/143 (83%) | Major facilitator superfamily protein | chr4:9777938-9779738 REVERSE LENGTH=544 |
