Probe CUST_50344_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50344_PI426222305 | JHI_St_60k_v1 | DMT400045837 | AACTCATGGATGACAGTCTAGTAGATTGGATTTTACGTGCCTTGGATGGCGATTCATCTT |
All Microarray Probes Designed to Gene DMG400017779
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50344_PI426222305 | JHI_St_60k_v1 | DMT400045837 | AACTCATGGATGACAGTCTAGTAGATTGGATTTTACGTGCCTTGGATGGCGATTCATCTT |
| CUST_50341_PI426222305 | JHI_St_60k_v1 | DMT400045838 | CGGACTCTGCAAAAATGTTGTCGTAGTATATGTAGGTGACATTTTTAAAGAGTTTGAAAG |
Microarray Signals from CUST_50344_PI426222305
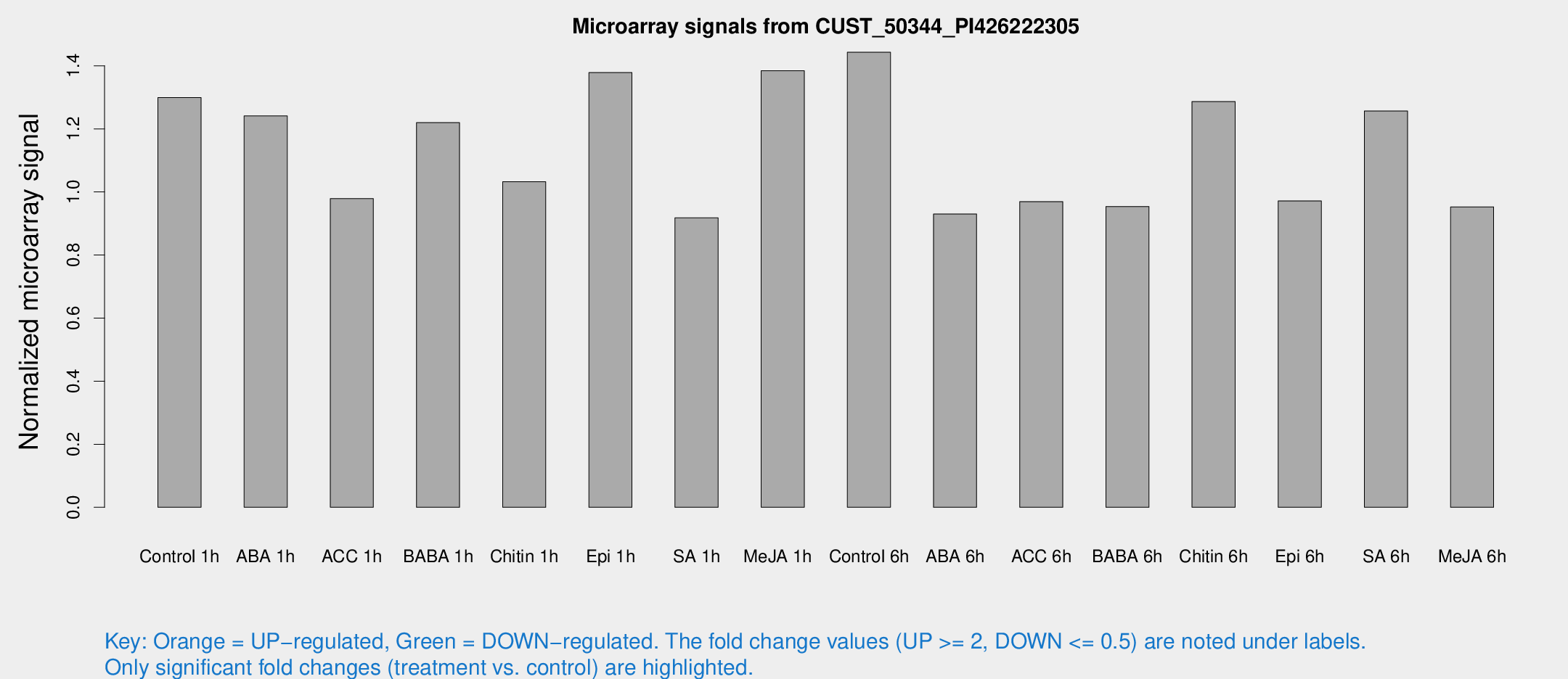
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 9.65865 | 3.55373 | 1.2993 | 0.608599 |
| ABA 1h | 7.72169 | 3.42636 | 1.24097 | 0.614104 |
| ACC 1h | 6.6349 | 3.85752 | 0.978892 | 0.567343 |
| BABA 1h | 8.02943 | 3.72454 | 1.21972 | 0.609975 |
| Chitin 1h | 6.12716 | 3.5588 | 1.0325 | 0.597927 |
| Epi 1h | 8.37318 | 3.52891 | 1.37849 | 0.664868 |
| SA 1h | 6.20602 | 3.49169 | 0.917845 | 0.516332 |
| Me-JA 1h | 7.71168 | 3.60061 | 1.38434 | 0.69331 |
| Control 6h | 11.4092 | 5.0921 | 1.443 | 0.666689 |
| ABA 6h | 6.51441 | 3.77875 | 0.930018 | 0.53868 |
| ACC 6h | 7.53441 | 4.49971 | 0.969246 | 0.561261 |
| BABA 6h | 7.04324 | 4.08157 | 0.953877 | 0.551126 |
| Chitin 6h | 9.88655 | 3.99998 | 1.28654 | 0.617544 |
| Epi 6h | 7.25738 | 4.23235 | 0.97148 | 0.56258 |
| SA 6h | 8.57212 | 3.81934 | 1.25662 | 0.617621 |
| Me-JA 6h | 6.24559 | 3.62415 | 0.952522 | 0.552251 |
Source Transcript PGSC0003DMT400045837 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34440.1 | +1 | 4e-141 | 431 | 247/467 (53%) | Protein kinase superfamily protein | chr4:16466008-16468748 FORWARD LENGTH=670 |
