Probe CUST_50339_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50339_PI426222305 | JHI_St_60k_v1 | DMT400045865 | ATGAATGAGCAAGCCAAGGAAAAGTATGAAGATGCCAAAGAGAAGGCCTCAGATGCTGCT |
All Microarray Probes Designed to Gene DMG400017792
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50339_PI426222305 | JHI_St_60k_v1 | DMT400045865 | ATGAATGAGCAAGCCAAGGAAAAGTATGAAGATGCCAAAGAGAAGGCCTCAGATGCTGCT |
Microarray Signals from CUST_50339_PI426222305
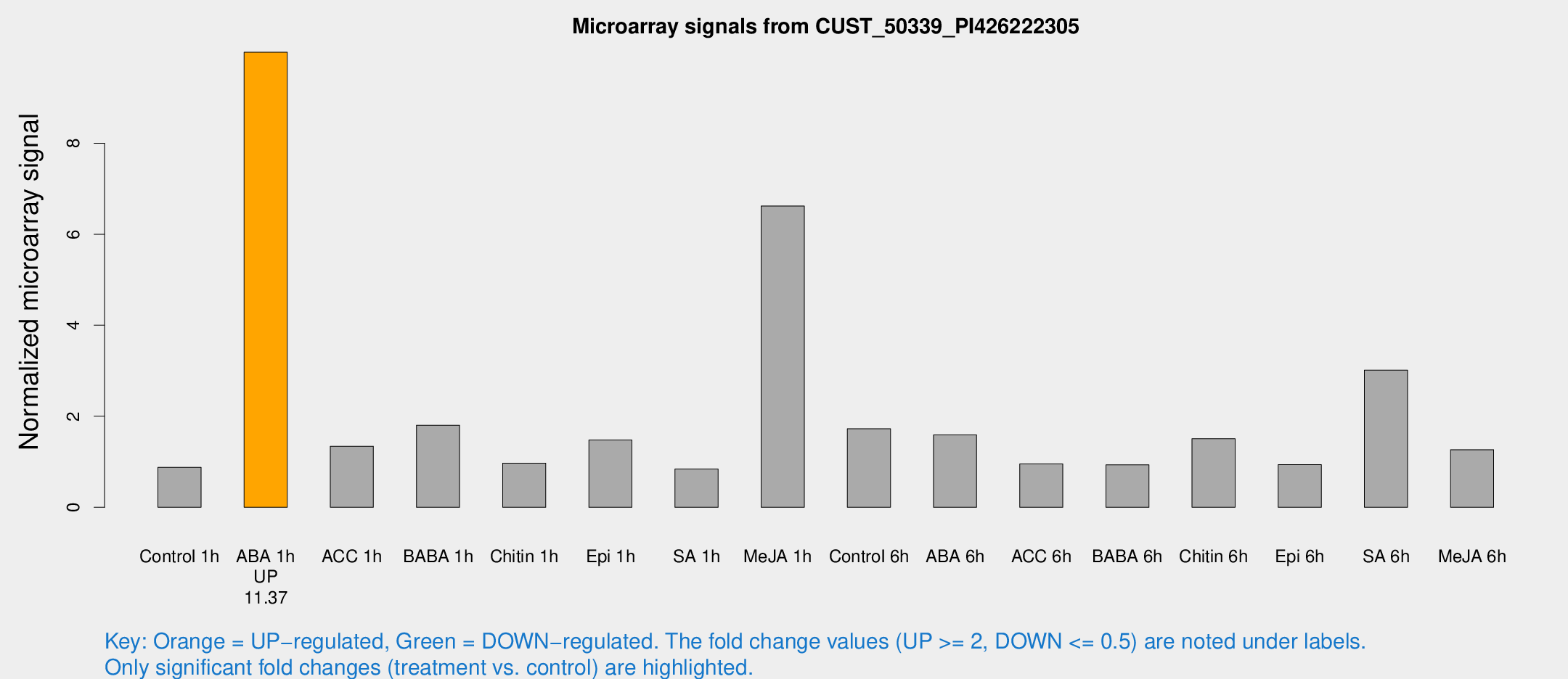
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.88545 | 3.42472 | 0.879059 | 0.509238 |
| ABA 1h | 60.7838 | 10.6135 | 9.99866 | 2.55442 |
| ACC 1h | 10.8595 | 4.67703 | 1.34009 | 0.76162 |
| BABA 1h | 12.3004 | 3.85292 | 1.8042 | 0.675668 |
| Chitin 1h | 5.8539 | 3.40874 | 0.971211 | 0.562417 |
| Epi 1h | 8.74289 | 3.37722 | 1.47768 | 0.617697 |
| SA 1h | 5.79312 | 3.37133 | 0.843034 | 0.48896 |
| Me-JA 1h | 66.9965 | 34.643 | 6.62095 | 16.2126 |
| Control 6h | 16.0982 | 9.48616 | 1.72538 | 1.15263 |
| ABA 6h | 12.7004 | 4.34905 | 1.59167 | 0.624939 |
| ACC 6h | 7.45728 | 4.42248 | 0.953918 | 0.552781 |
| BABA 6h | 6.98277 | 4.06671 | 0.932174 | 0.540578 |
| Chitin 6h | 12.0083 | 4.31058 | 1.50668 | 0.67782 |
| Epi 6h | 7.14584 | 4.20176 | 0.936768 | 0.542599 |
| SA 6h | 25.0515 | 9.92087 | 3.01451 | 2.53785 |
| Me-JA 6h | 9.04616 | 3.68838 | 1.2657 | 0.604729 |
Source Transcript PGSC0003DMT400045865 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G18340.1 | +1 | 4e-40 | 150 | 142/418 (34%) | late embryogenesis abundant domain-containing protein / LEA domain-containing protein | chr2:7969400-7971025 FORWARD LENGTH=456 |
