Probe CUST_50045_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50045_PI426222305 | JHI_St_60k_v1 | DMT400065416 | TACCATGTTGGAGGAAAGGCAAAGAAAGCAGCTGATAGGGTGATTCCAGAGTTAGCATAG |
All Microarray Probes Designed to Gene DMG400025433
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_50054_PI426222305 | JHI_St_60k_v1 | DMT400065417 | GTATGAACTTCCTAATGACAAGTTTCCTTTTCAACAAAGTTACTTTCGCCGTCAGAATTT |
| CUST_50045_PI426222305 | JHI_St_60k_v1 | DMT400065416 | TACCATGTTGGAGGAAAGGCAAAGAAAGCAGCTGATAGGGTGATTCCAGAGTTAGCATAG |
Microarray Signals from CUST_50045_PI426222305
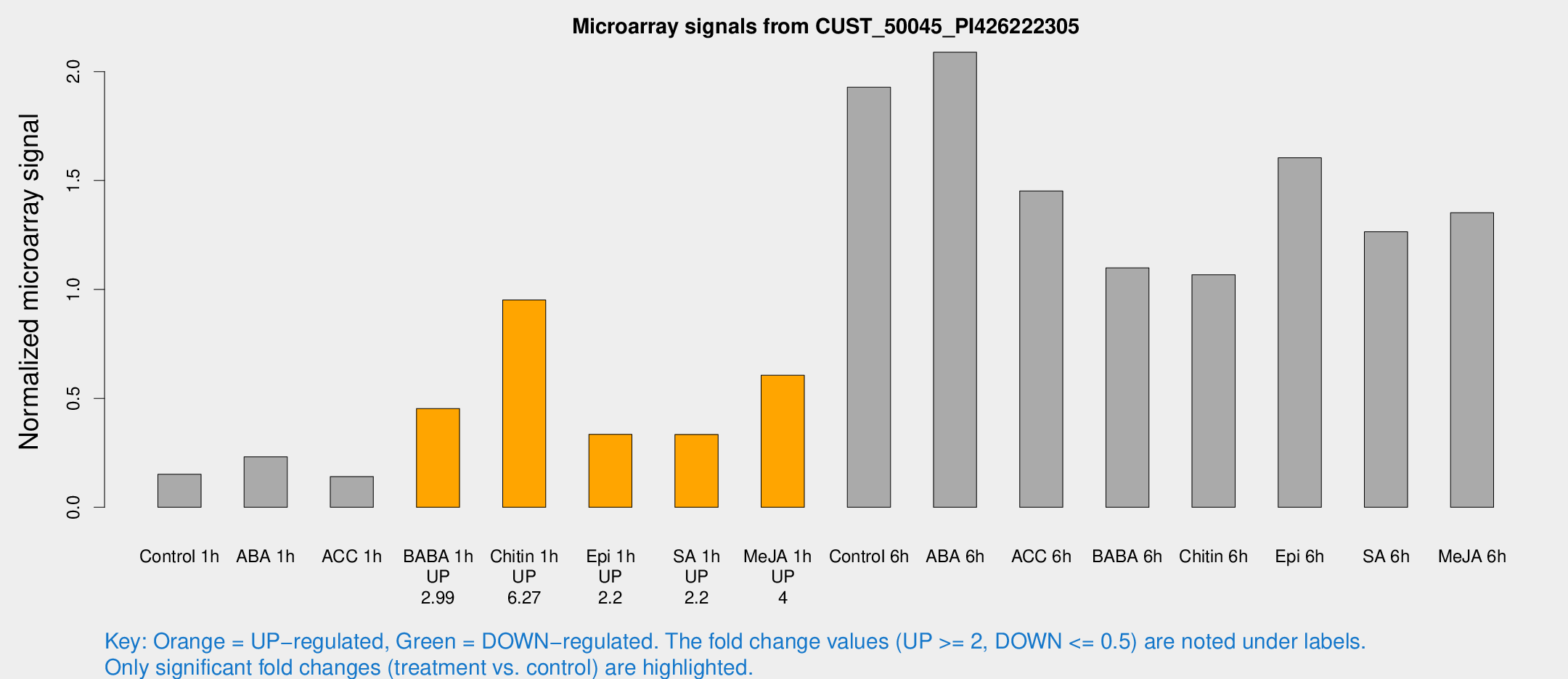
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 262.775 | 38.8718 | 0.151731 | 0.030372 |
| ABA 1h | 359.56 | 67.7867 | 0.231825 | 0.0306368 |
| ACC 1h | 248.5 | 27.1467 | 0.140819 | 0.0243337 |
| BABA 1h | 847.642 | 266.334 | 0.453431 | 0.139646 |
| Chitin 1h | 1459.06 | 107.038 | 0.951655 | 0.0550007 |
| Epi 1h | 511.36 | 96.3141 | 0.334548 | 0.0640277 |
| SA 1h | 624.8 | 174.332 | 0.33419 | 0.0705752 |
| Me-JA 1h | 875.053 | 180.642 | 0.606201 | 0.071161 |
| Control 6h | 3369.95 | 522.536 | 1.92818 | 0.148296 |
| ABA 6h | 3904.11 | 734.36 | 2.08894 | 0.282925 |
| ACC 6h | 2830.3 | 163.722 | 1.45222 | 0.159533 |
| BABA 6h | 2095.15 | 133.714 | 1.09877 | 0.0634786 |
| Chitin 6h | 1936.96 | 138.979 | 1.06722 | 0.0616615 |
| Epi 6h | 3181.38 | 610.868 | 1.60448 | 0.269467 |
| SA 6h | 2233.36 | 457.105 | 1.26449 | 0.149319 |
| Me-JA 6h | 2361.38 | 407.336 | 1.35235 | 0.13999 |
Source Transcript PGSC0003DMT400065416 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G52450.1 | +1 | 3e-175 | 504 | 260/406 (64%) | MATE efflux family protein | chr5:21289042-21291749 REVERSE LENGTH=486 |
