Probe CUST_49592_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49592_PI426222305 | JHI_St_60k_v1 | DMT400013634 | TTCAAGCCATGCTTTTGTCGTTCAAGATGCACTTTTCGTCAATTAACCAGGATTGAGGCG |
All Microarray Probes Designed to Gene DMG400005330
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49592_PI426222305 | JHI_St_60k_v1 | DMT400013634 | TTCAAGCCATGCTTTTGTCGTTCAAGATGCACTTTTCGTCAATTAACCAGGATTGAGGCG |
Microarray Signals from CUST_49592_PI426222305
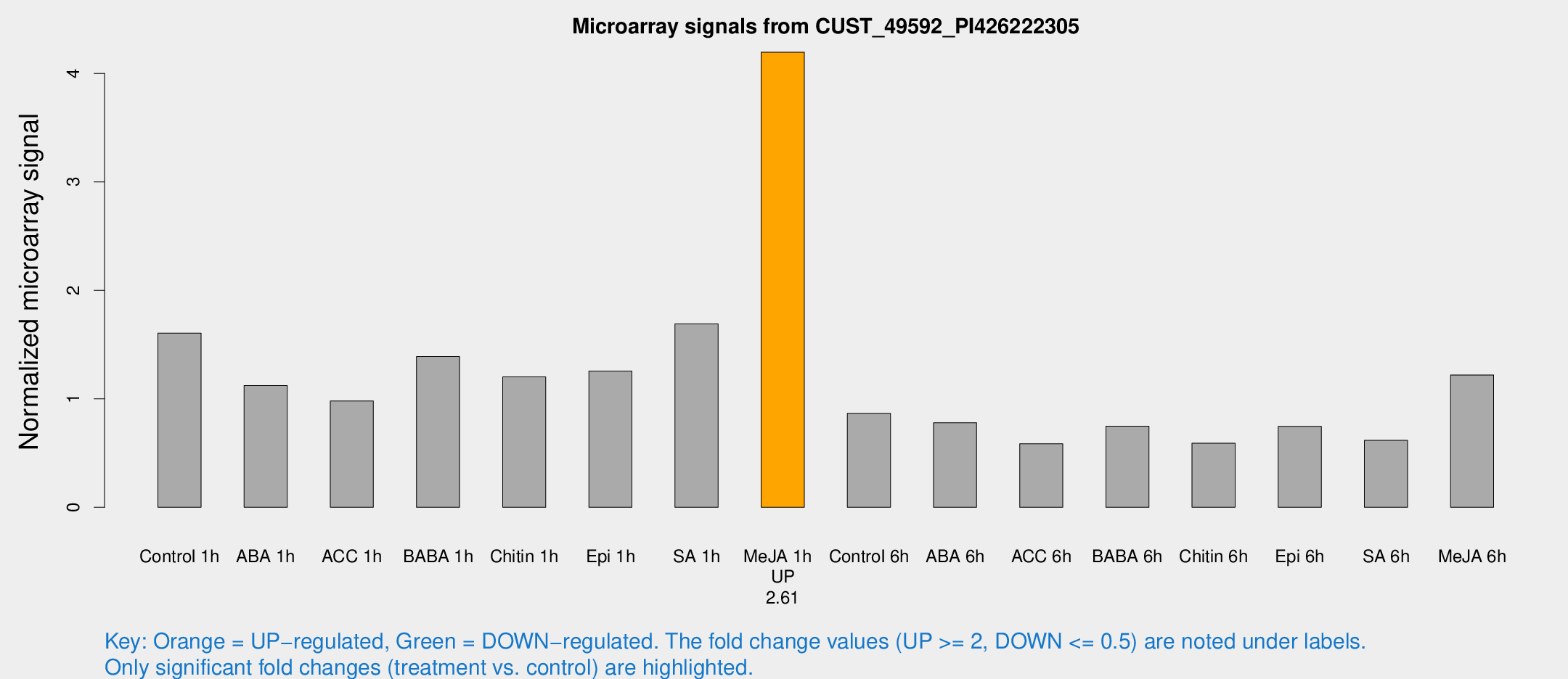
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 166.225 | 25.4513 | 1.60552 | 0.223241 |
| ABA 1h | 104.882 | 21.5772 | 1.1226 | 0.162012 |
| ACC 1h | 115.546 | 37.7785 | 0.980577 | 0.319285 |
| BABA 1h | 141.435 | 27.7377 | 1.38919 | 0.162054 |
| Chitin 1h | 109.71 | 7.21343 | 1.20207 | 0.112874 |
| Epi 1h | 113.062 | 17.8944 | 1.25642 | 0.196648 |
| SA 1h | 176.531 | 13.1241 | 1.69117 | 0.170897 |
| Me-JA 1h | 348.046 | 26.2274 | 4.195 | 0.245323 |
| Control 6h | 93.6647 | 22.1546 | 0.866448 | 0.149714 |
| ABA 6h | 84.094 | 5.96822 | 0.779127 | 0.055277 |
| ACC 6h | 69.1212 | 9.47259 | 0.585722 | 0.0473446 |
| BABA 6h | 86.4049 | 12.4434 | 0.74735 | 0.081575 |
| Chitin 6h | 65.6413 | 11.7926 | 0.591305 | 0.101272 |
| Epi 6h | 87.6039 | 15.2967 | 0.745915 | 0.112144 |
| SA 6h | 73.6197 | 24.8693 | 0.617348 | 0.232646 |
| Me-JA 6h | 128.908 | 26.8894 | 1.21946 | 0.176909 |
Source Transcript PGSC0003DMT400013634 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G63220.2 | +2 | 8e-171 | 494 | 231/349 (66%) | Galactose oxidase/kelch repeat superfamily protein | chr3:23357540-23358598 REVERSE LENGTH=352 |
