Probe CUST_49345_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49345_PI426222305 | JHI_St_60k_v1 | DMT400056192 | CAAATTGTATGTTGGTTGCTTTAGCTCAGAAAGCCAGCAACATCTTGTTTCTATTGAAAA |
All Microarray Probes Designed to Gene DMG400021831
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49345_PI426222305 | JHI_St_60k_v1 | DMT400056192 | CAAATTGTATGTTGGTTGCTTTAGCTCAGAAAGCCAGCAACATCTTGTTTCTATTGAAAA |
Microarray Signals from CUST_49345_PI426222305
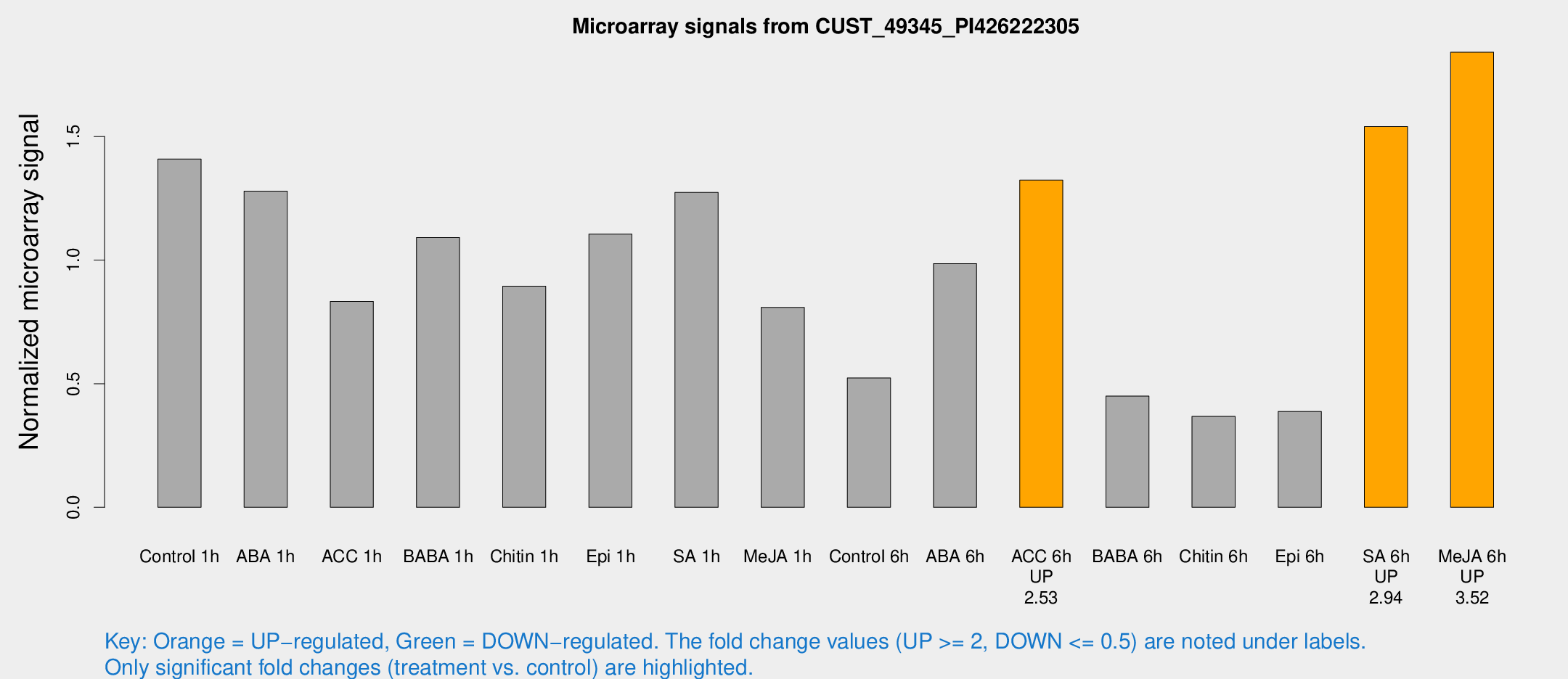
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3506.29 | 455.706 | 1.40882 | 0.0994708 |
| ABA 1h | 2830.58 | 418.968 | 1.27911 | 0.186119 |
| ACC 1h | 2286.86 | 621.059 | 0.832698 | 0.207078 |
| BABA 1h | 2820.09 | 746.907 | 1.09144 | 0.240897 |
| Chitin 1h | 1997.39 | 239.929 | 0.894384 | 0.0684898 |
| Epi 1h | 2371.3 | 245.901 | 1.10539 | 0.108923 |
| SA 1h | 3296.27 | 527.464 | 1.27352 | 0.142661 |
| Me-JA 1h | 1652.36 | 236.062 | 0.808378 | 0.0538097 |
| Control 6h | 1314.88 | 189.251 | 0.523199 | 0.0363415 |
| ABA 6h | 2714.87 | 583.506 | 0.98566 | 0.200902 |
| ACC 6h | 3731.12 | 216.134 | 1.32347 | 0.257684 |
| BABA 6h | 1235.24 | 71.5116 | 0.449848 | 0.0260091 |
| Chitin 6h | 963.432 | 58.5033 | 0.368032 | 0.0278634 |
| Epi 6h | 1077.32 | 84.3802 | 0.387516 | 0.0562734 |
| SA 6h | 3828.85 | 636.596 | 1.53974 | 0.261445 |
| Me-JA 6h | 4517.74 | 352.385 | 1.84084 | 0.156156 |
Source Transcript PGSC0003DMT400056192 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G22240.1 | +2 | 0.0 | 935 | 454/510 (89%) | myo-inositol-1-phosphate synthase 2 | chr2:9451901-9453938 REVERSE LENGTH=510 |
