Probe CUST_4930_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_4930_PI426222305 | JHI_St_60k_v1 | DMT400038572 | GGGTTTCTCTCCTTGGCTTGGAAAGATAATCCCCTTTTGACTGTTTCTTCATGGCACTAA |
All Microarray Probes Designed to Gene DMG400014900
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_5261_PI426222305 | JHI_St_60k_v1 | DMT400038573 | CTTGTTCTGATCCTTTAGTTTTCTAGGTTTTGTTTCTTTTTTGGTGTCATTAGGAGGAGA |
| CUST_4930_PI426222305 | JHI_St_60k_v1 | DMT400038572 | GGGTTTCTCTCCTTGGCTTGGAAAGATAATCCCCTTTTGACTGTTTCTTCATGGCACTAA |
Microarray Signals from CUST_4930_PI426222305
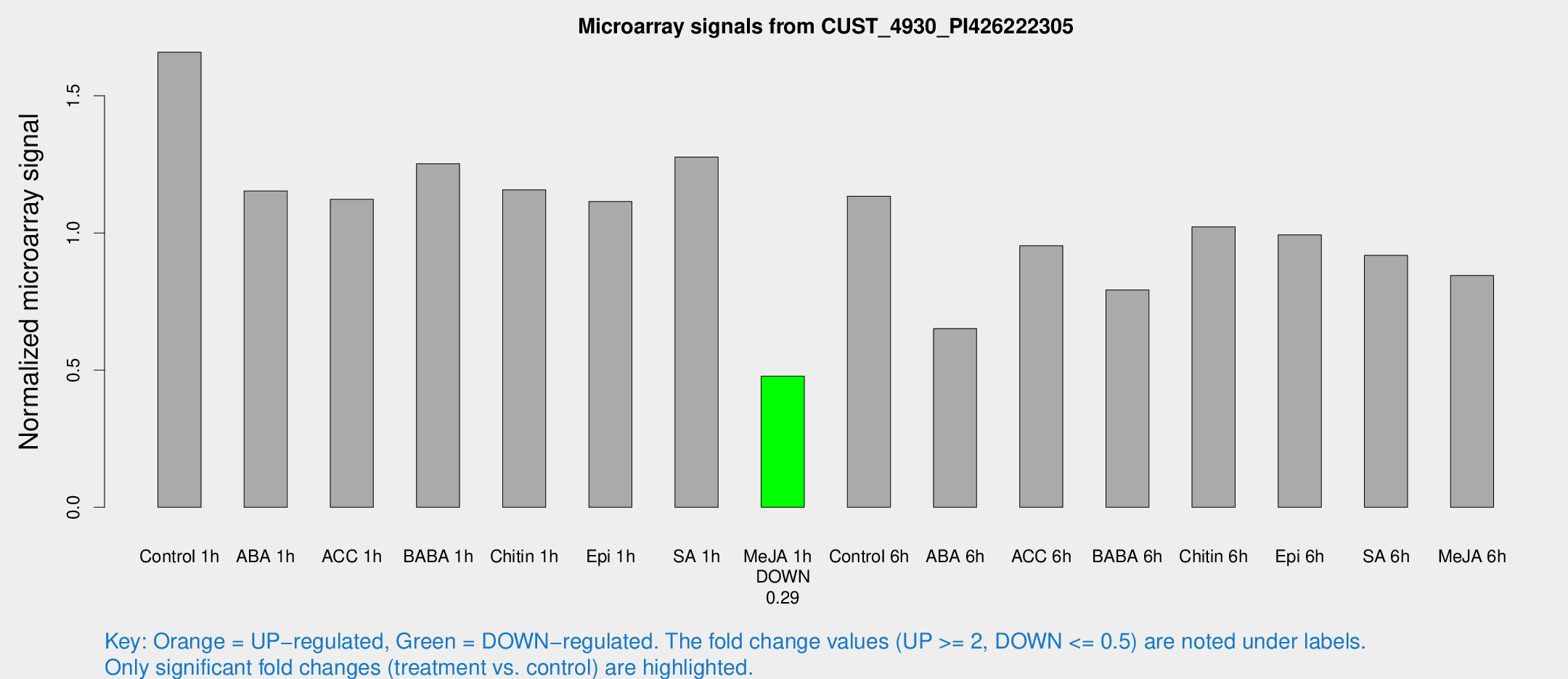
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8811.63 | 1243.81 | 1.65873 | 0.168094 |
| ABA 1h | 5350.8 | 543.785 | 1.1531 | 0.0733687 |
| ACC 1h | 6231.22 | 1147.31 | 1.1226 | 0.144299 |
| BABA 1h | 6809.75 | 1795.67 | 1.25245 | 0.257019 |
| Chitin 1h | 5512.71 | 737.743 | 1.15725 | 0.0856168 |
| Epi 1h | 5011.54 | 289.433 | 1.11447 | 0.0643478 |
| SA 1h | 7153.63 | 1431.08 | 1.2769 | 0.233168 |
| Me-JA 1h | 2058.43 | 239.287 | 0.478398 | 0.0276314 |
| Control 6h | 6227.46 | 1410.25 | 1.13397 | 0.192685 |
| ABA 6h | 3606.82 | 208.803 | 0.651428 | 0.0376153 |
| ACC 6h | 5717.65 | 477.985 | 0.953135 | 0.0550329 |
| BABA 6h | 4691.21 | 634.402 | 0.792437 | 0.119115 |
| Chitin 6h | 5677.33 | 367.076 | 1.02247 | 0.0590359 |
| Epi 6h | 5914.07 | 758.541 | 0.992922 | 0.0999847 |
| SA 6h | 4944.98 | 977.238 | 0.9182 | 0.0986339 |
| Me-JA 6h | 4419.32 | 468.296 | 0.845227 | 0.0488035 |
Source Transcript PGSC0003DMT400038572 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G66770.1 | +1 | 1e-177 | 515 | 278/488 (57%) | GRAS family transcription factor | chr5:26660723-26662477 FORWARD LENGTH=584 |
