Probe CUST_49285_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49285_PI426222305 | JHI_St_60k_v1 | DMT400059103 | GCTTGTAGGTTGTATAGAAAAATTGAGGTGTTCATGGTAAAAGTGTCTGGATTTCTAGTT |
All Microarray Probes Designed to Gene DMG400022956
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_49267_PI426222305 | JHI_St_60k_v1 | DMT400059101 | GCTTGTAGGTTGTATAGAAAAATTGAGGTGTTCATGGTAAAAGTGTCTGGATTTCTAGTT |
| CUST_49285_PI426222305 | JHI_St_60k_v1 | DMT400059103 | GCTTGTAGGTTGTATAGAAAAATTGAGGTGTTCATGGTAAAAGTGTCTGGATTTCTAGTT |
Microarray Signals from CUST_49285_PI426222305
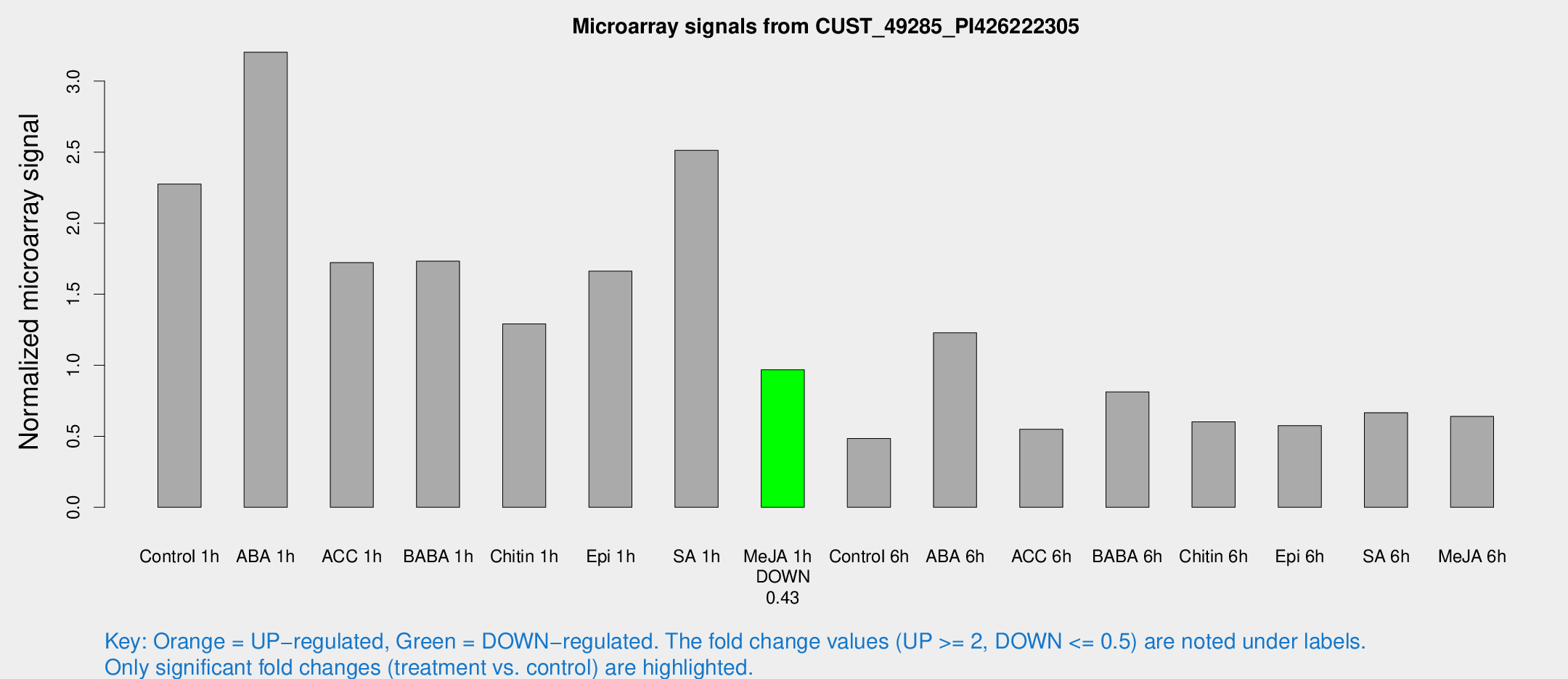
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3440.03 | 512.632 | 2.27528 | 0.210889 |
| ABA 1h | 4287.4 | 591.615 | 3.20356 | 0.299772 |
| ACC 1h | 2893.95 | 783.503 | 1.72207 | 0.471548 |
| BABA 1h | 2700.56 | 702.635 | 1.73245 | 0.375074 |
| Chitin 1h | 1730.43 | 137.295 | 1.29151 | 0.0746068 |
| Epi 1h | 2133.14 | 123.309 | 1.66263 | 0.0960259 |
| SA 1h | 3954.72 | 753.666 | 2.51322 | 0.315286 |
| Me-JA 1h | 1185.17 | 132.141 | 0.968655 | 0.0559952 |
| Control 6h | 761.735 | 174.048 | 0.484447 | 0.0839542 |
| ABA 6h | 2012.36 | 392.471 | 1.22813 | 0.1832 |
| ACC 6h | 944.824 | 109.085 | 0.548755 | 0.0317705 |
| BABA 6h | 1382.57 | 210.728 | 0.812914 | 0.101118 |
| Chitin 6h | 956.308 | 87.8193 | 0.601515 | 0.0674717 |
| Epi 6h | 974.108 | 111.146 | 0.575449 | 0.0477726 |
| SA 6h | 1031.69 | 226.653 | 0.666061 | 0.0907589 |
| Me-JA 6h | 998.891 | 237.156 | 0.639722 | 0.10127 |
Source Transcript PGSC0003DMT400059103 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G05190.1 | +2 | 4e-20 | 96 | 44/87 (51%) | Protein of unknown function (DUF3133) | chr5:1541853-1543875 FORWARD LENGTH=615 |
