Probe CUST_48823_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48823_PI426222305 | JHI_St_60k_v1 | DMT400056101 | GATCATGGGCTTGATAGATTTATGGAGATTTACTTTATGGGTGTGTTTGGTATGAAAGAA |
All Microarray Probes Designed to Gene DMG400021791
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48826_PI426222305 | JHI_St_60k_v1 | DMT400056100 | GATCATGGGCTTGATAGATTTATGGAGATTTACTTTATGGGTGTGTTTGGTATGAAAGAA |
| CUST_48823_PI426222305 | JHI_St_60k_v1 | DMT400056101 | GATCATGGGCTTGATAGATTTATGGAGATTTACTTTATGGGTGTGTTTGGTATGAAAGAA |
Microarray Signals from CUST_48823_PI426222305
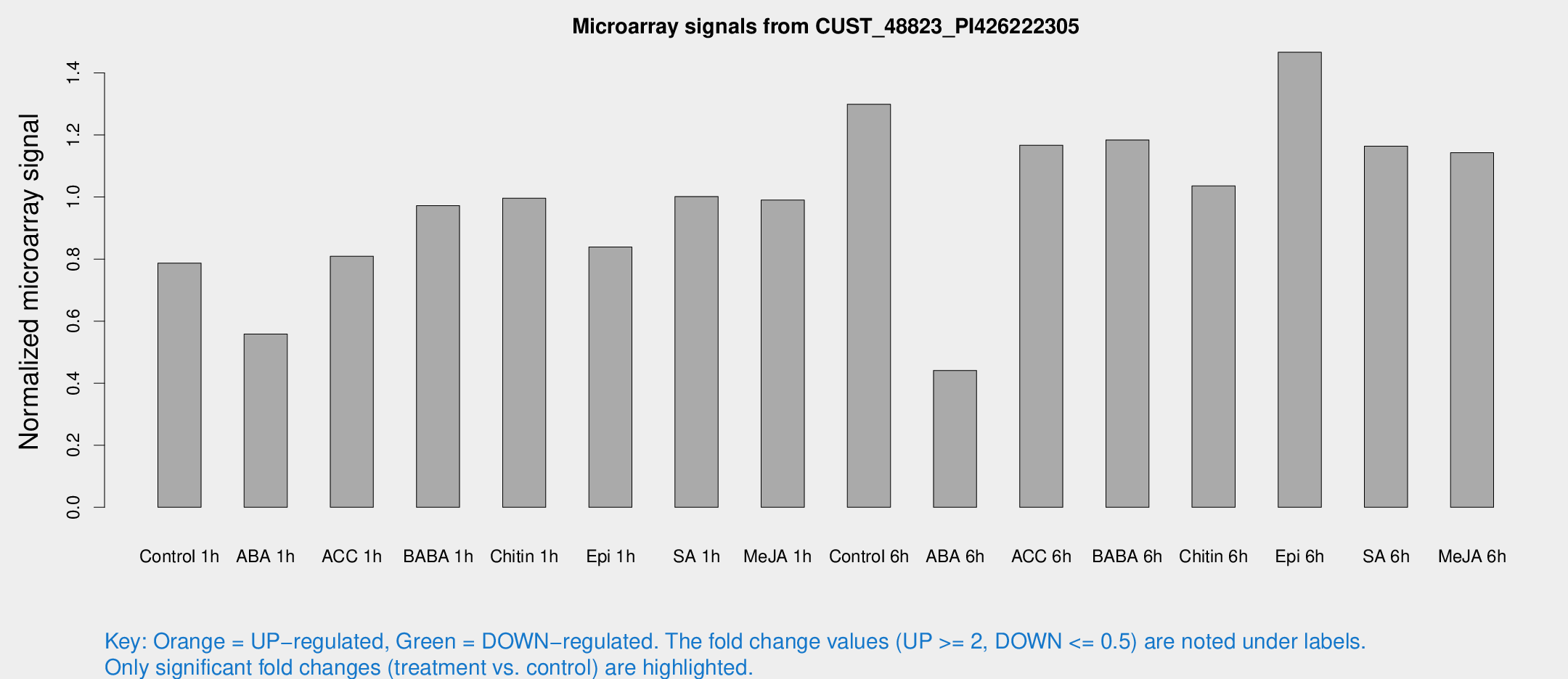
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 3233.49 | 528.78 | 0.787212 | 0.124932 |
| ABA 1h | 1972.78 | 113.949 | 0.558479 | 0.0412764 |
| ACC 1h | 3399.63 | 533.737 | 0.808903 | 0.0879007 |
| BABA 1h | 3885.5 | 701.577 | 0.972245 | 0.0948742 |
| Chitin 1h | 3633.55 | 439.841 | 0.996237 | 0.0920835 |
| Epi 1h | 2909.01 | 171.44 | 0.838873 | 0.054476 |
| SA 1h | 4311.47 | 871.65 | 1.00166 | 0.188909 |
| Me-JA 1h | 3336.58 | 562.973 | 0.99026 | 0.128222 |
| Control 6h | 5469.39 | 1123.6 | 1.29918 | 0.181957 |
| ABA 6h | 1928.15 | 339.902 | 0.441117 | 0.0510283 |
| ACC 6h | 5365.26 | 316.402 | 1.16709 | 0.149348 |
| BABA 6h | 5304.83 | 306.995 | 1.18405 | 0.0888807 |
| Chitin 6h | 4402.7 | 254.221 | 1.03579 | 0.0598079 |
| Epi 6h | 6724.84 | 940.459 | 1.46669 | 0.220316 |
| SA 6h | 4990.4 | 1247.73 | 1.16372 | 0.221004 |
| Me-JA 6h | 4580.13 | 414.496 | 1.14269 | 0.0659786 |
Source Transcript PGSC0003DMT400056101 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G05050.1 | +1 | 1e-27 | 108 | 72/147 (49%) | ubiquitin 11 | chr4:2588271-2588960 REVERSE LENGTH=229 |
