Probe CUST_48699_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48699_PI426222305 | JHI_St_60k_v1 | DMT400080295 | CCTGTAACATATATGCAGTACTCCTAGAAACAGCTGTACAGTTAAGTACTAATATGTATG |
All Microarray Probes Designed to Gene DMG400031261
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48698_PI426222305 | JHI_St_60k_v1 | DMT400080296 | CCCTTAGTTTCGAACAACTAGTTTTACTTCCAAAATTATCATGCTTCTGATGCATCACTA |
| CUST_48699_PI426222305 | JHI_St_60k_v1 | DMT400080295 | CCTGTAACATATATGCAGTACTCCTAGAAACAGCTGTACAGTTAAGTACTAATATGTATG |
Microarray Signals from CUST_48699_PI426222305
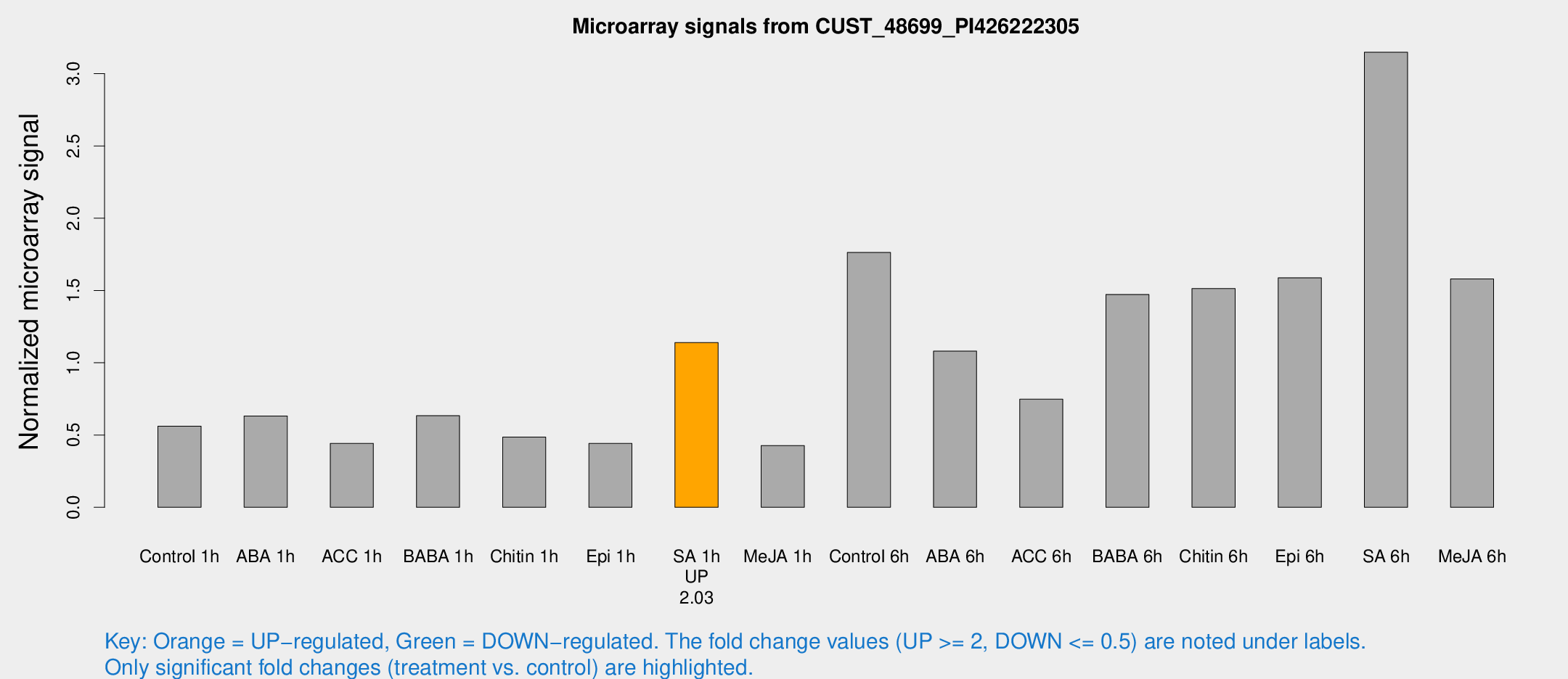
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 126.948 | 15.8484 | 0.560952 | 0.0476431 |
| ABA 1h | 126.705 | 16.4792 | 0.632101 | 0.0835389 |
| ACC 1h | 127.199 | 49.4359 | 0.441767 | 0.235824 |
| BABA 1h | 145.979 | 35.2842 | 0.633683 | 0.112092 |
| Chitin 1h | 101.891 | 21.8252 | 0.485739 | 0.0652451 |
| Epi 1h | 88.1809 | 15.1142 | 0.441545 | 0.0787621 |
| SA 1h | 263.644 | 25.2917 | 1.13992 | 0.0672917 |
| Me-JA 1h | 78.8005 | 9.19543 | 0.427234 | 0.0303649 |
| Control 6h | 408.322 | 71.6159 | 1.76279 | 0.176751 |
| ABA 6h | 257.563 | 18.7952 | 1.08061 | 0.0761499 |
| ACC 6h | 205.054 | 53.1166 | 0.748002 | 0.222814 |
| BABA 6h | 369.309 | 25.6587 | 1.47249 | 0.0862784 |
| Chitin 6h | 359.542 | 21.0886 | 1.51347 | 0.0887046 |
| Epi 6h | 406.194 | 54.6685 | 1.58778 | 0.100289 |
| SA 6h | 707.913 | 95.4726 | 3.14846 | 0.18251 |
| Me-JA 6h | 412.424 | 173.405 | 1.57958 | 0.540741 |
Source Transcript PGSC0003DMT400080295 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14790.1 | +2 | 0.0 | 1284 | 683/1109 (62%) | RNA-dependent RNA polymerase 1 | chr1:5094317-5097817 REVERSE LENGTH=1107 |
