Probe CUST_48698_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48698_PI426222305 | JHI_St_60k_v1 | DMT400080296 | CCCTTAGTTTCGAACAACTAGTTTTACTTCCAAAATTATCATGCTTCTGATGCATCACTA |
All Microarray Probes Designed to Gene DMG400031261
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48698_PI426222305 | JHI_St_60k_v1 | DMT400080296 | CCCTTAGTTTCGAACAACTAGTTTTACTTCCAAAATTATCATGCTTCTGATGCATCACTA |
| CUST_48699_PI426222305 | JHI_St_60k_v1 | DMT400080295 | CCTGTAACATATATGCAGTACTCCTAGAAACAGCTGTACAGTTAAGTACTAATATGTATG |
Microarray Signals from CUST_48698_PI426222305
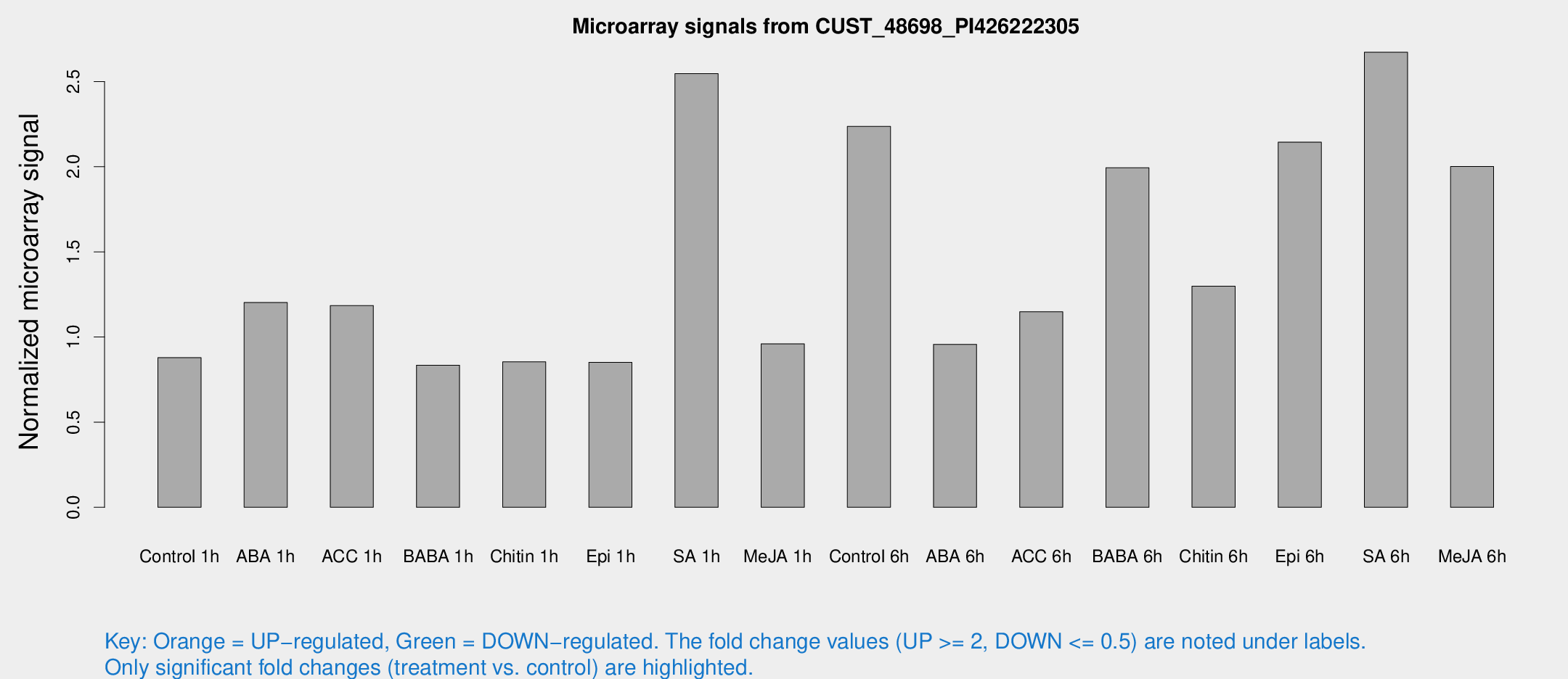
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.42845 | 3.63991 | 0.878838 | 0.448644 |
| ABA 1h | 9.71811 | 3.58483 | 1.20236 | 0.530562 |
| ACC 1h | 10.906 | 4.07254 | 1.18414 | 0.520772 |
| BABA 1h | 6.70837 | 3.90182 | 0.834539 | 0.483747 |
| Chitin 1h | 6.41785 | 3.74166 | 0.853906 | 0.495276 |
| Epi 1h | 6.14377 | 3.57228 | 0.851025 | 0.4932 |
| SA 1h | 22.2129 | 3.87433 | 2.54666 | 0.531882 |
| Me-JA 1h | 6.54726 | 3.81453 | 0.959239 | 0.555419 |
| Control 6h | 21.0906 | 6.23493 | 2.23687 | 0.677519 |
| ABA 6h | 8.74967 | 4.07958 | 0.956984 | 0.479088 |
| ACC 6h | 12.3346 | 4.67706 | 1.14834 | 0.538343 |
| BABA 6h | 22.5171 | 7.86608 | 1.99362 | 1.05137 |
| Chitin 6h | 13.4713 | 5.64361 | 1.29826 | 0.60869 |
| Epi 6h | 22.8172 | 8.56824 | 2.14401 | 0.800772 |
| SA 6h | 26.776 | 12.367 | 2.672 | 1.87211 |
| Me-JA 6h | 27.4878 | 18.9611 | 2.00075 | 1.71563 |
Source Transcript PGSC0003DMT400080296 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G14790.1 | +2 | 0.0 | 949 | 519/873 (59%) | RNA-dependent RNA polymerase 1 | chr1:5094317-5097817 REVERSE LENGTH=1107 |
