Probe CUST_48349_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48349_PI426222305 | JHI_St_60k_v1 | DMT400043223 | GAAAGAGAAAGATCTCTACTTAATCGCCAAAACGAATCAAAGTTTGGATATCATGCTCTA |
All Microarray Probes Designed to Gene DMG400016776
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48315_PI426222305 | JHI_St_60k_v1 | DMT400043224 | CTCTAAAGGGAGAAAATGAGCAAAGAATAAGCTTTTCCTCAACTTCTTTTAGTTTGGTGT |
| CUST_48349_PI426222305 | JHI_St_60k_v1 | DMT400043223 | GAAAGAGAAAGATCTCTACTTAATCGCCAAAACGAATCAAAGTTTGGATATCATGCTCTA |
Microarray Signals from CUST_48349_PI426222305
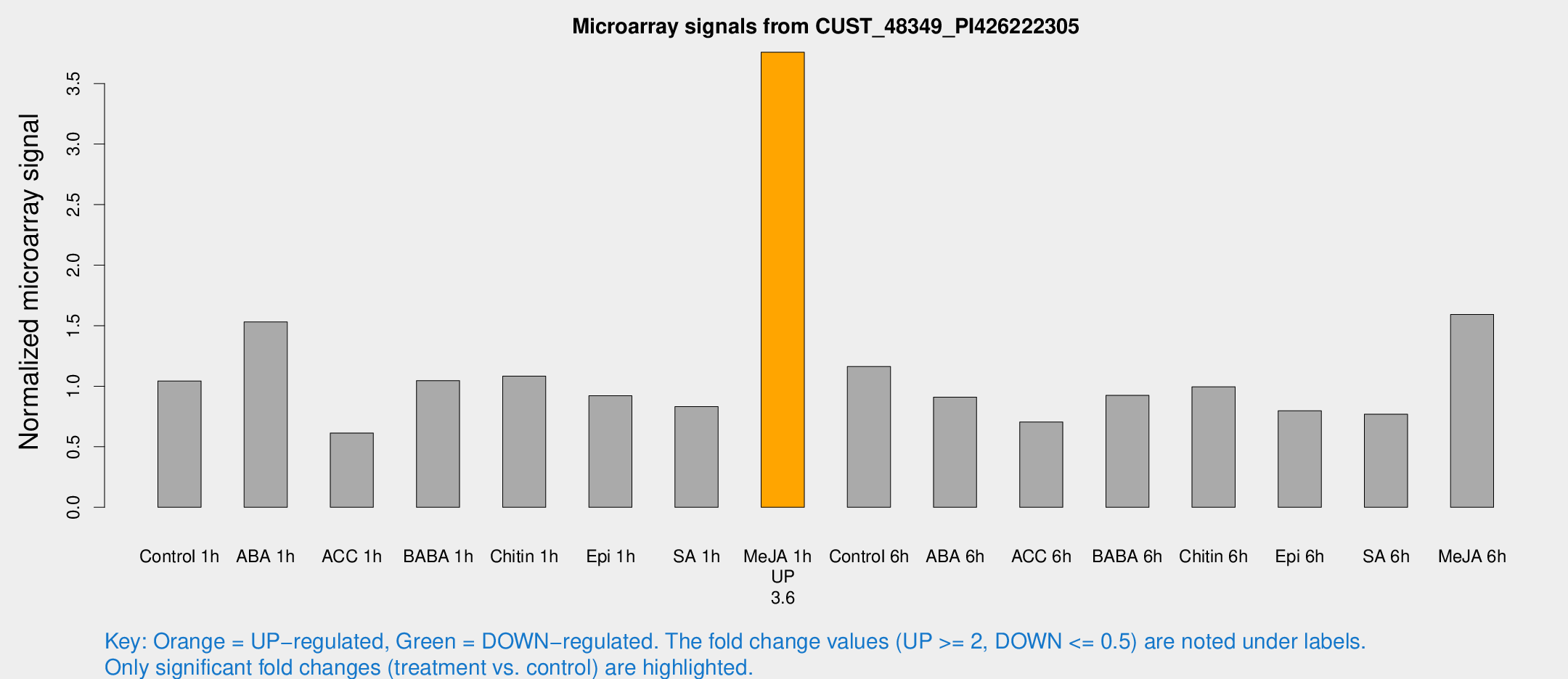
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 207.252 | 37.6103 | 1.04371 | 0.121412 |
| ABA 1h | 277.235 | 66.8649 | 1.53065 | 0.282502 |
| ACC 1h | 142.686 | 47.3812 | 0.613784 | 0.245698 |
| BABA 1h | 198.838 | 28.5241 | 1.0463 | 0.062816 |
| Chitin 1h | 188.794 | 15.0162 | 1.08286 | 0.0646505 |
| Epi 1h | 155.154 | 15.0522 | 0.921244 | 0.0759984 |
| SA 1h | 171.333 | 35.567 | 0.831477 | 0.162269 |
| Me-JA 1h | 594.217 | 43.3085 | 3.75856 | 0.217809 |
| Control 6h | 243.767 | 62.4517 | 1.1632 | 0.256909 |
| ABA 6h | 188.797 | 20.4699 | 0.910007 | 0.115505 |
| ACC 6h | 159.018 | 22.8119 | 0.705186 | 0.0436635 |
| BABA 6h | 206.508 | 40.3796 | 0.924263 | 0.164104 |
| Chitin 6h | 204.244 | 12.2482 | 0.995776 | 0.0597113 |
| Epi 6h | 182.023 | 38.0969 | 0.797375 | 0.259538 |
| SA 6h | 148.143 | 15.0774 | 0.768847 | 0.0475963 |
| Me-JA 6h | 320.716 | 68.8379 | 1.59248 | 0.222661 |
Source Transcript PGSC0003DMT400043223 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26210.1 | +1 | 9e-90 | 283 | 156/400 (39%) | cytochrome P450, family 71, subfamily B, polypeptide 23 | chr3:9593329-9595202 REVERSE LENGTH=501 |
