Probe CUST_48315_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48315_PI426222305 | JHI_St_60k_v1 | DMT400043224 | CTCTAAAGGGAGAAAATGAGCAAAGAATAAGCTTTTCCTCAACTTCTTTTAGTTTGGTGT |
All Microarray Probes Designed to Gene DMG400016776
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48315_PI426222305 | JHI_St_60k_v1 | DMT400043224 | CTCTAAAGGGAGAAAATGAGCAAAGAATAAGCTTTTCCTCAACTTCTTTTAGTTTGGTGT |
| CUST_48349_PI426222305 | JHI_St_60k_v1 | DMT400043223 | GAAAGAGAAAGATCTCTACTTAATCGCCAAAACGAATCAAAGTTTGGATATCATGCTCTA |
Microarray Signals from CUST_48315_PI426222305
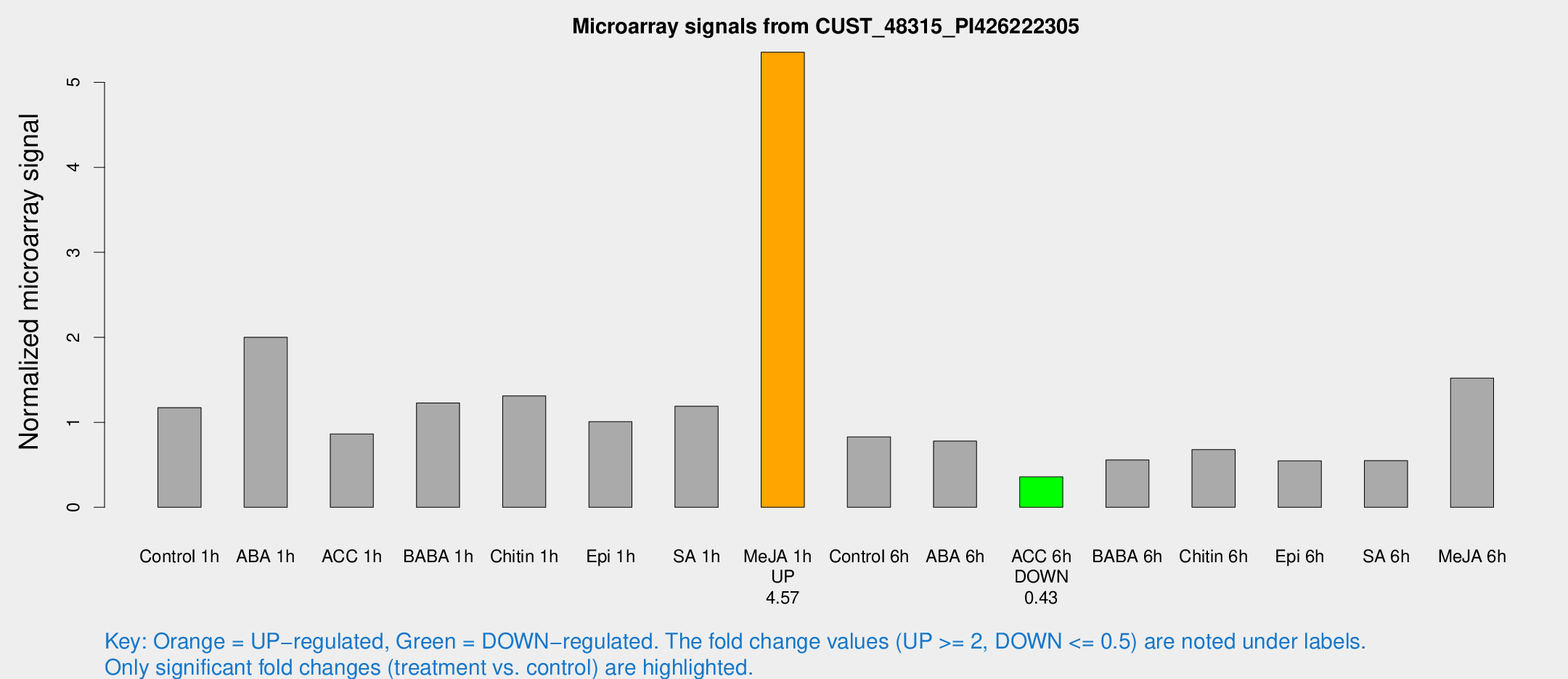
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 250.637 | 53.1284 | 1.17254 | 0.163934 |
| ABA 1h | 386.117 | 94.7706 | 2.00021 | 0.371947 |
| ACC 1h | 190.907 | 42.242 | 0.862277 | 0.148187 |
| BABA 1h | 253.335 | 49.2005 | 1.22739 | 0.140346 |
| Chitin 1h | 243.679 | 21.2343 | 1.31152 | 0.0772623 |
| Epi 1h | 181.198 | 19.8684 | 1.00752 | 0.102796 |
| SA 1h | 256.05 | 39.1938 | 1.18856 | 0.169988 |
| Me-JA 1h | 901.284 | 67.6427 | 5.35425 | 0.309613 |
| Control 6h | 183.388 | 45.4625 | 0.828212 | 0.161223 |
| ABA 6h | 172.861 | 22.7502 | 0.778884 | 0.0921258 |
| ACC 6h | 87.5247 | 17.5984 | 0.358526 | 0.0253867 |
| BABA 6h | 129.097 | 12.4981 | 0.557347 | 0.0485832 |
| Chitin 6h | 149.337 | 12.634 | 0.679184 | 0.0723551 |
| Epi 6h | 131.169 | 24.1804 | 0.546896 | 0.156023 |
| SA 6h | 115.082 | 20.082 | 0.548112 | 0.0423124 |
| Me-JA 6h | 326.223 | 71.4886 | 1.51969 | 0.216482 |
Source Transcript PGSC0003DMT400043224 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G26210.1 | +2 | 1e-114 | 355 | 195/486 (40%) | cytochrome P450, family 71, subfamily B, polypeptide 23 | chr3:9593329-9595202 REVERSE LENGTH=501 |
