Probe CUST_48216_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48216_PI426222305 | JHI_St_60k_v1 | DMT400055910 | GCAATGAATGTTCTTGGATTTGATATTAATGACACCAGTGCCACTGCCTCTAAAGGAGCT |
All Microarray Probes Designed to Gene DMG400021719
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_48216_PI426222305 | JHI_St_60k_v1 | DMT400055910 | GCAATGAATGTTCTTGGATTTGATATTAATGACACCAGTGCCACTGCCTCTAAAGGAGCT |
| CUST_48228_PI426222305 | JHI_St_60k_v1 | DMT400055911 | ACACGAAAAAGAAACCAACATAGTCAGGGATGAGGCAATTTTGTACTTCTTTTAGTTTCC |
Microarray Signals from CUST_48216_PI426222305
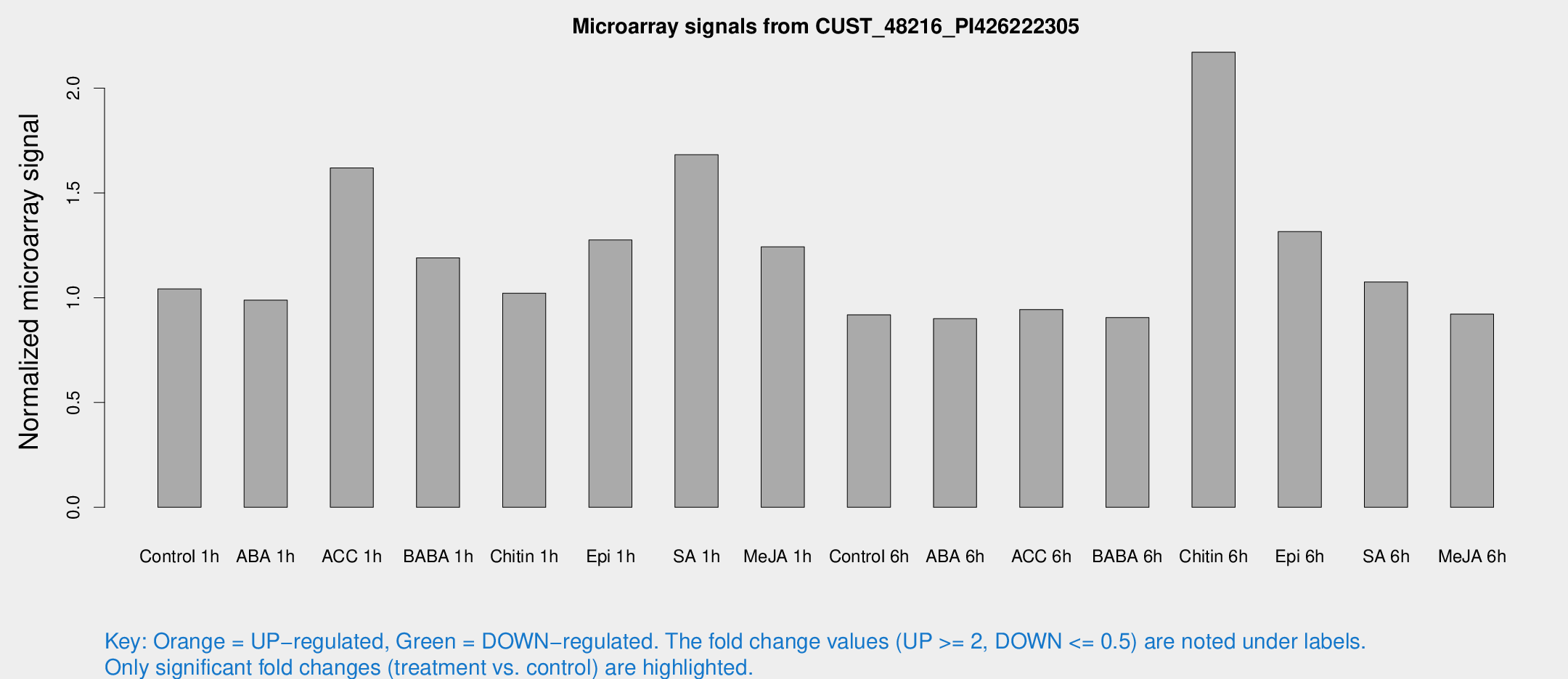
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.35982 | 3.74579 | 1.04247 | 0.542594 |
| ABA 1h | 6.13641 | 3.56573 | 0.988578 | 0.572479 |
| ACC 1h | 13.1963 | 4.49213 | 1.61918 | 0.658061 |
| BABA 1h | 8.34371 | 3.84708 | 1.19028 | 0.593592 |
| Chitin 1h | 6.44056 | 3.73933 | 1.02165 | 0.591635 |
| Epi 1h | 8.23779 | 3.60072 | 1.27552 | 0.626496 |
| SA 1h | 13.6706 | 4.48029 | 1.68216 | 0.825896 |
| Me-JA 1h | 7.13436 | 3.72709 | 1.24265 | 0.657661 |
| Control 6h | 6.43586 | 3.72896 | 0.918236 | 0.531995 |
| ABA 6h | 6.72437 | 3.82862 | 0.900047 | 0.511956 |
| ACC 6h | 7.73505 | 4.58138 | 0.943821 | 0.546543 |
| BABA 6h | 7.10756 | 4.11057 | 0.905268 | 0.521441 |
| Chitin 6h | 23.5309 | 14.3251 | 2.1713 | 1.61071 |
| Epi 6h | 10.4681 | 4.59327 | 1.31577 | 0.589556 |
| SA 6h | 7.49384 | 4.05557 | 1.07535 | 0.587476 |
| Me-JA 6h | 6.4288 | 3.72926 | 0.921563 | 0.533653 |
Source Transcript PGSC0003DMT400055910 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G21330.1 | +1 | 2e-37 | 134 | 85/178 (48%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr4:11349922-11350694 FORWARD LENGTH=207 |
