Probe CUST_47563_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47563_PI426222305 | JHI_St_60k_v1 | DMT400027239 | CACCAACGTCTGCTCCATTTCGACATTAGAATGATTTCTAGTAATAAATTTGAAAGACGA |
All Microarray Probes Designed to Gene DMG400010504
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47563_PI426222305 | JHI_St_60k_v1 | DMT400027239 | CACCAACGTCTGCTCCATTTCGACATTAGAATGATTTCTAGTAATAAATTTGAAAGACGA |
Microarray Signals from CUST_47563_PI426222305
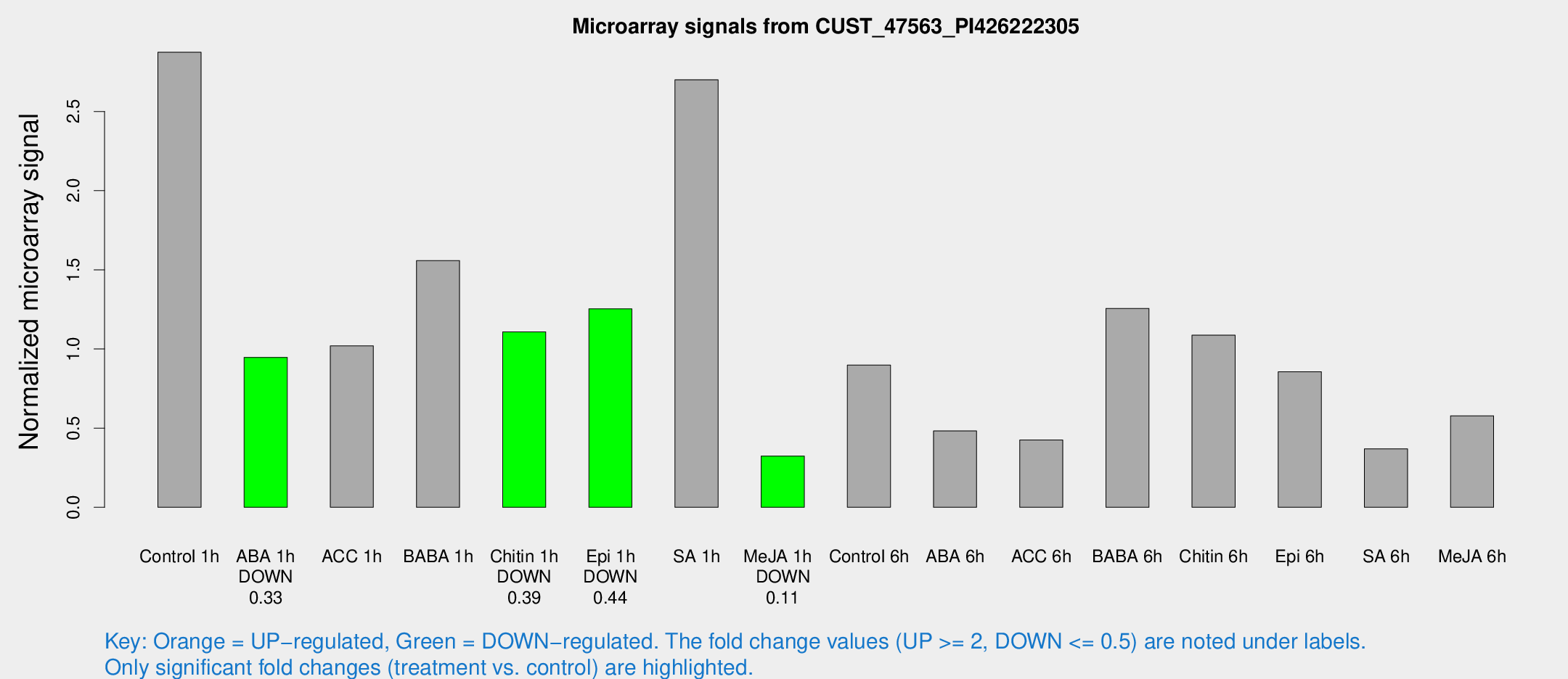
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 144.95 | 43.9053 | 2.87412 | 0.658327 |
| ABA 1h | 39.3743 | 4.61285 | 0.946992 | 0.096261 |
| ACC 1h | 72.7003 | 31.9898 | 1.01951 | 1.00302 |
| BABA 1h | 81.2494 | 26.7859 | 1.55812 | 0.544329 |
| Chitin 1h | 47.5303 | 7.69941 | 1.10838 | 0.191986 |
| Epi 1h | 52.4111 | 9.77632 | 1.25369 | 0.254859 |
| SA 1h | 132.558 | 23.1894 | 2.70007 | 0.311269 |
| Me-JA 1h | 13.5952 | 4.61925 | 0.32393 | 0.0994943 |
| Control 6h | 44.1412 | 9.42495 | 0.898613 | 0.142899 |
| ABA 6h | 25.3838 | 6.48962 | 0.482078 | 0.101336 |
| ACC 6h | 25.8595 | 8.05965 | 0.425035 | 0.162159 |
| BABA 6h | 65.5115 | 5.35619 | 1.25623 | 0.102908 |
| Chitin 6h | 54.3859 | 6.72361 | 1.08696 | 0.101514 |
| Epi 6h | 57.7757 | 21.9163 | 0.85647 | 0.533527 |
| SA 6h | 23.0078 | 12.032 | 0.36936 | 0.206489 |
| Me-JA 6h | 33.5237 | 15.8114 | 0.578204 | 0.25852 |
Source Transcript PGSC0003DMT400027239 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G66980.1 | +3 | 3e-73 | 162 | 78/181 (43%) | suppressor of npr1-1 constitutive 4 | chr1:24997491-25001961 REVERSE LENGTH=1118 |
