Probe CUST_47124_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47124_PI426222305 | JHI_St_60k_v1 | DMT400030422 | TAGTTCGAGCCAGAAGCCGTGAATTTATCATTTGAATTTCTATGGTGTTCATCATATTTG |
All Microarray Probes Designed to Gene DMG400011651
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_47124_PI426222305 | JHI_St_60k_v1 | DMT400030422 | TAGTTCGAGCCAGAAGCCGTGAATTTATCATTTGAATTTCTATGGTGTTCATCATATTTG |
| CUST_47127_PI426222305 | JHI_St_60k_v1 | DMT400030421 | GAGGAAAGTGAAGAAACTATTGTTGTCCTTTATTGGGATTCGTGCTTCTAGGAATATAAA |
Microarray Signals from CUST_47124_PI426222305
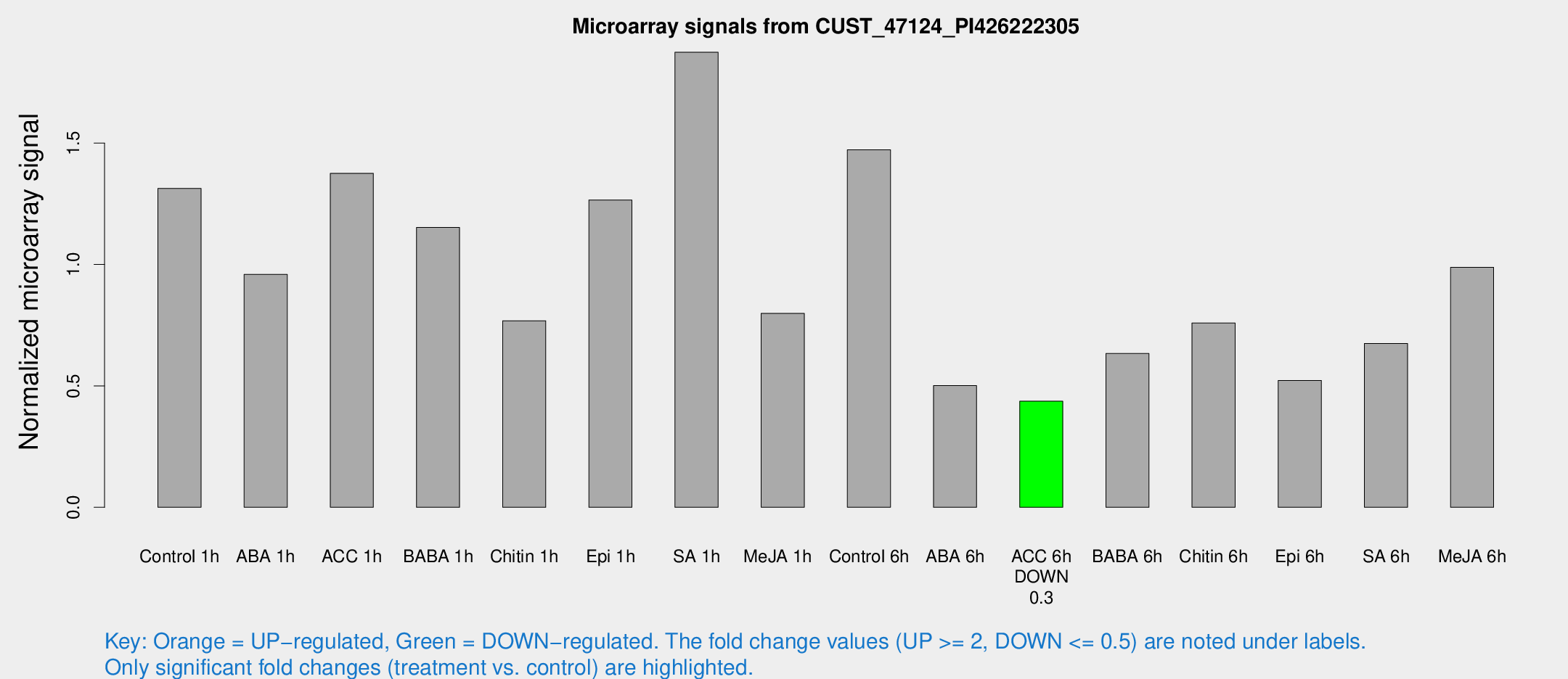
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 24.7456 | 10.5896 | 1.31357 | 0.908491 |
| ABA 1h | 13.9206 | 4.45269 | 0.959523 | 0.399365 |
| ACC 1h | 23.8565 | 8.21449 | 1.37537 | 0.616498 |
| BABA 1h | 19.9625 | 8.13404 | 1.15262 | 0.597919 |
| Chitin 1h | 10.2824 | 3.06151 | 0.768405 | 0.274417 |
| Epi 1h | 15.5351 | 3.09247 | 1.26564 | 0.257112 |
| SA 1h | 27.6571 | 3.97583 | 1.87421 | 0.245933 |
| Me-JA 1h | 10.2202 | 3.40262 | 0.798627 | 0.300048 |
| Control 6h | 20.9841 | 3.31882 | 1.47221 | 0.241311 |
| ABA 6h | 8.37602 | 3.2469 | 0.501121 | 0.238415 |
| ACC 6h | 7.09675 | 3.74484 | 0.436954 | 0.223851 |
| BABA 6h | 11.1499 | 3.74502 | 0.633366 | 0.25003 |
| Chitin 6h | 12.5194 | 3.52734 | 0.758936 | 0.273301 |
| Epi 6h | 8.99797 | 3.66038 | 0.522639 | 0.24904 |
| SA 6h | 11.928 | 6.0799 | 0.674522 | 0.327779 |
| Me-JA 6h | 14.8983 | 3.71105 | 0.98806 | 0.256536 |
Source Transcript PGSC0003DMT400030422 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
