Probe CUST_45926_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45926_PI426222305 | JHI_St_60k_v1 | DMT400047955 | TACTAGGACTATTCATCTTTTAGAATTTCATTTTCACCGTGAAGTTGGGATCCGTGTCAC |
All Microarray Probes Designed to Gene DMG400018636
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45926_PI426222305 | JHI_St_60k_v1 | DMT400047955 | TACTAGGACTATTCATCTTTTAGAATTTCATTTTCACCGTGAAGTTGGGATCCGTGTCAC |
| CUST_45891_PI426222305 | JHI_St_60k_v1 | DMT400047950 | TTGAAATACTCTCATCCTAGCCCCAAGCATCGAGAGAGGAATAACAGAGCTGATCCTTTA |
Microarray Signals from CUST_45926_PI426222305
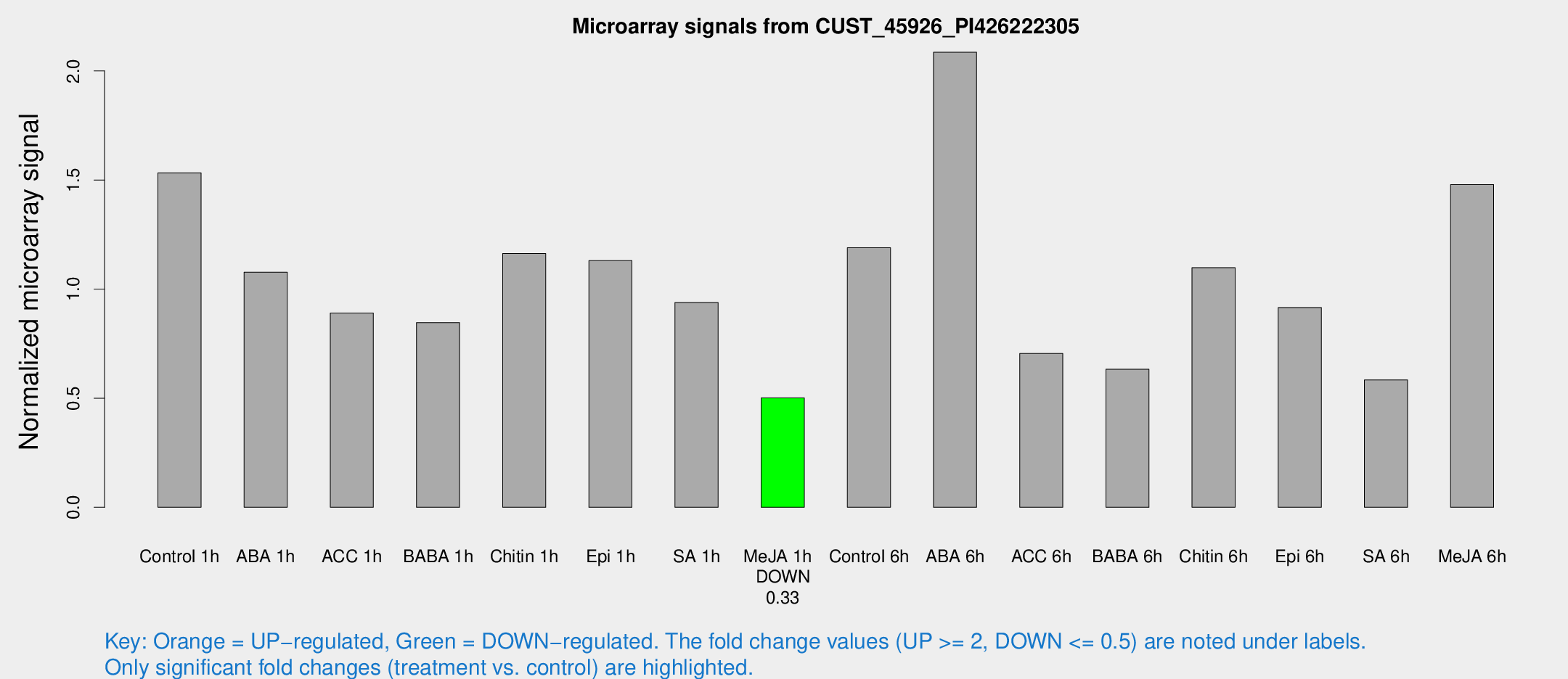
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 25.6379 | 3.95735 | 1.53319 | 0.237005 |
| ABA 1h | 19.455 | 8.23721 | 1.07774 | 0.746536 |
| ACC 1h | 15.854 | 4.95027 | 0.890398 | 0.330607 |
| BABA 1h | 15.0819 | 5.325 | 0.84687 | 0.321555 |
| Chitin 1h | 17.882 | 3.95436 | 1.16336 | 0.340195 |
| Epi 1h | 17.1096 | 4.5529 | 1.13046 | 0.325789 |
| SA 1h | 18.2593 | 5.8942 | 0.938797 | 0.515735 |
| Me-JA 1h | 6.73446 | 3.7871 | 0.501217 | 0.278996 |
| Control 6h | 20.8178 | 4.73771 | 1.19004 | 0.285997 |
| ABA 6h | 38.971 | 9.25592 | 2.08594 | 0.516521 |
| ACC 6h | 13.8479 | 4.5599 | 0.704907 | 0.24119 |
| BABA 6h | 13.4755 | 5.44486 | 0.632875 | 0.280965 |
| Chitin 6h | 20.2246 | 4.45656 | 1.09844 | 0.292451 |
| Epi 6h | 19.7589 | 7.9469 | 0.915883 | 0.370072 |
| SA 6h | 10.3172 | 4.1289 | 0.583937 | 0.279088 |
| Me-JA 6h | 26.0174 | 7.51471 | 1.47897 | 0.298659 |
Source Transcript PGSC0003DMT400047955 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G60620.1 | +3 | 1e-68 | 238 | 110/134 (82%) | glycerol-3-phosphate acyltransferase 9 | chr5:24367266-24369647 FORWARD LENGTH=376 |
