Probe CUST_45891_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45891_PI426222305 | JHI_St_60k_v1 | DMT400047950 | TTGAAATACTCTCATCCTAGCCCCAAGCATCGAGAGAGGAATAACAGAGCTGATCCTTTA |
All Microarray Probes Designed to Gene DMG400018636
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45926_PI426222305 | JHI_St_60k_v1 | DMT400047955 | TACTAGGACTATTCATCTTTTAGAATTTCATTTTCACCGTGAAGTTGGGATCCGTGTCAC |
| CUST_45891_PI426222305 | JHI_St_60k_v1 | DMT400047950 | TTGAAATACTCTCATCCTAGCCCCAAGCATCGAGAGAGGAATAACAGAGCTGATCCTTTA |
Microarray Signals from CUST_45891_PI426222305
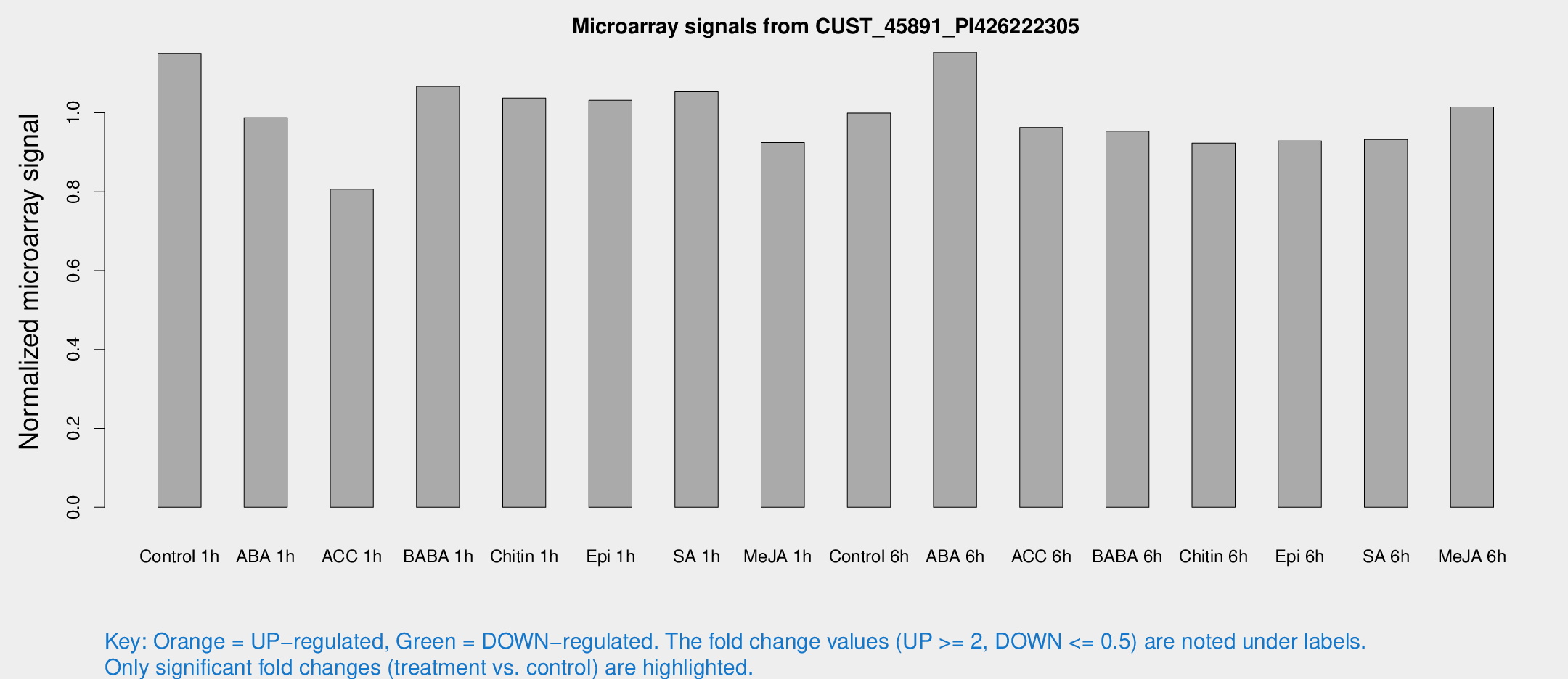
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1244 | 98.9416 | 1.15061 | 0.06649 |
| ABA 1h | 948.699 | 95.2656 | 0.987874 | 0.0571193 |
| ACC 1h | 945.642 | 209 | 0.806747 | 0.146818 |
| BABA 1h | 1176.37 | 271.922 | 1.06705 | 0.172258 |
| Chitin 1h | 1011.83 | 100.625 | 1.03729 | 0.0599676 |
| Epi 1h | 969.64 | 100.226 | 1.03159 | 0.107892 |
| SA 1h | 1205.96 | 211.804 | 1.05349 | 0.158294 |
| Me-JA 1h | 822.648 | 98.5725 | 0.92449 | 0.0534931 |
| Control 6h | 1139.37 | 267.875 | 0.999242 | 0.179196 |
| ABA 6h | 1333.16 | 137.4 | 1.15362 | 0.0722242 |
| ACC 6h | 1189.27 | 68.8892 | 0.963035 | 0.0932997 |
| BABA 6h | 1153.52 | 88.316 | 0.95353 | 0.067693 |
| Chitin 6h | 1065.66 | 97.5836 | 0.923303 | 0.069893 |
| Epi 6h | 1157.89 | 182.909 | 0.929015 | 0.109102 |
| SA 6h | 1119.65 | 364.447 | 0.932502 | 0.24853 |
| Me-JA 6h | 1092.42 | 81.1277 | 1.015 | 0.0586727 |
Source Transcript PGSC0003DMT400047950 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G60620.1 | +1 | 2e-61 | 197 | 90/107 (84%) | glycerol-3-phosphate acyltransferase 9 | chr5:24367266-24369647 FORWARD LENGTH=376 |
