Probe CUST_45820_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45820_PI426222305 | JHI_St_60k_v1 | DMT400050261 | CTGTTAGCCTTTGATTGTATGAGATCAGGTTGGTTGGCTTACTAATCTTTTATTGTGTTT |
All Microarray Probes Designed to Gene DMG400019523
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45820_PI426222305 | JHI_St_60k_v1 | DMT400050261 | CTGTTAGCCTTTGATTGTATGAGATCAGGTTGGTTGGCTTACTAATCTTTTATTGTGTTT |
| CUST_45843_PI426222305 | JHI_St_60k_v1 | DMT400050262 | CTGTTAGCCTTTGATTGTATGAGATCAGGTTGGTTGGCTTACTAATCTTTTATTGTGTTT |
Microarray Signals from CUST_45820_PI426222305
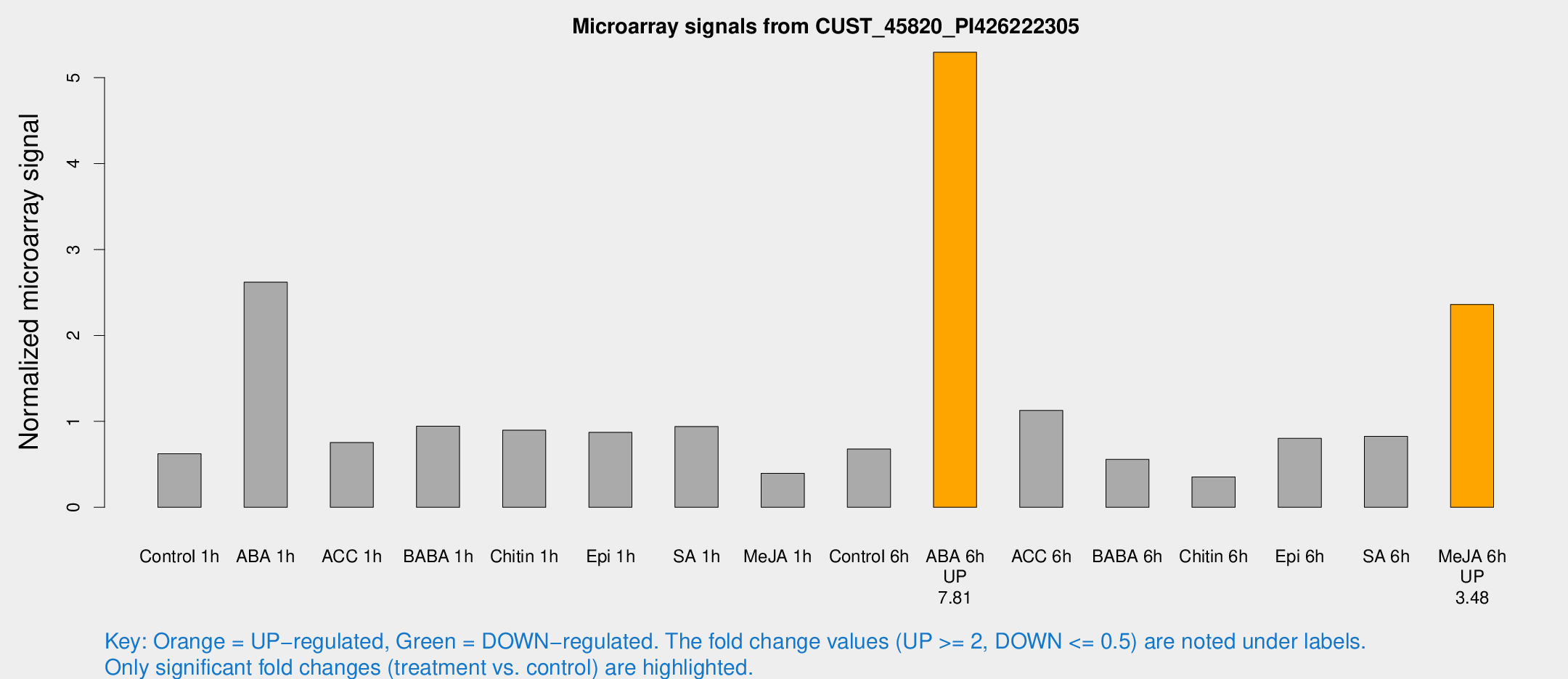
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18.0402 | 7.60686 | 0.62352 | 0.361238 |
| ABA 1h | 59.3063 | 15.415 | 2.62062 | 0.66188 |
| ACC 1h | 24.5457 | 12.8282 | 0.753646 | 0.432863 |
| BABA 1h | 27.7878 | 10.4322 | 0.943697 | 0.50472 |
| Chitin 1h | 22.3498 | 7.13456 | 0.896852 | 0.427929 |
| Epi 1h | 21.4829 | 8.85056 | 0.87166 | 0.419143 |
| SA 1h | 30.0566 | 12.2231 | 0.940148 | 0.620839 |
| Me-JA 1h | 8.054 | 3.28536 | 0.395305 | 0.172736 |
| Control 6h | 18.0272 | 5.73487 | 0.67828 | 0.258976 |
| ABA 6h | 140.39 | 26.3968 | 5.29576 | 0.932779 |
| ACC 6h | 32.8583 | 7.49297 | 1.1268 | 0.492376 |
| BABA 6h | 16.2659 | 4.92476 | 0.558487 | 0.154627 |
| Chitin 6h | 9.7943 | 3.61563 | 0.352421 | 0.155125 |
| Epi 6h | 23.358 | 5.77317 | 0.802082 | 0.183651 |
| SA 6h | 25.6203 | 10.3493 | 0.82653 | 0.459337 |
| Me-JA 6h | 58.1127 | 10.2546 | 2.36108 | 0.588965 |
Source Transcript PGSC0003DMT400050261 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G61430.1 | +3 | 6e-109 | 334 | 193/358 (54%) | NAC domain containing protein 100 | chr5:24701328-24702553 REVERSE LENGTH=336 |
