Probe CUST_45809_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45809_PI426222305 | JHI_St_60k_v1 | DMT400050269 | CACATAACCTGCCGGTTAGTTAATTTCTTGTTTAATTTTTATATGGTAGTGTGGCCTGTT |
All Microarray Probes Designed to Gene DMG400019526
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45797_PI426222305 | JHI_St_60k_v1 | DMT400050270 | TAACCTGCCGGTGTGGATCTCAATTTTGCTATATATGTGGAGAAACATGGAGCGAAAGCC |
| CUST_45809_PI426222305 | JHI_St_60k_v1 | DMT400050269 | CACATAACCTGCCGGTTAGTTAATTTCTTGTTTAATTTTTATATGGTAGTGTGGCCTGTT |
Microarray Signals from CUST_45809_PI426222305
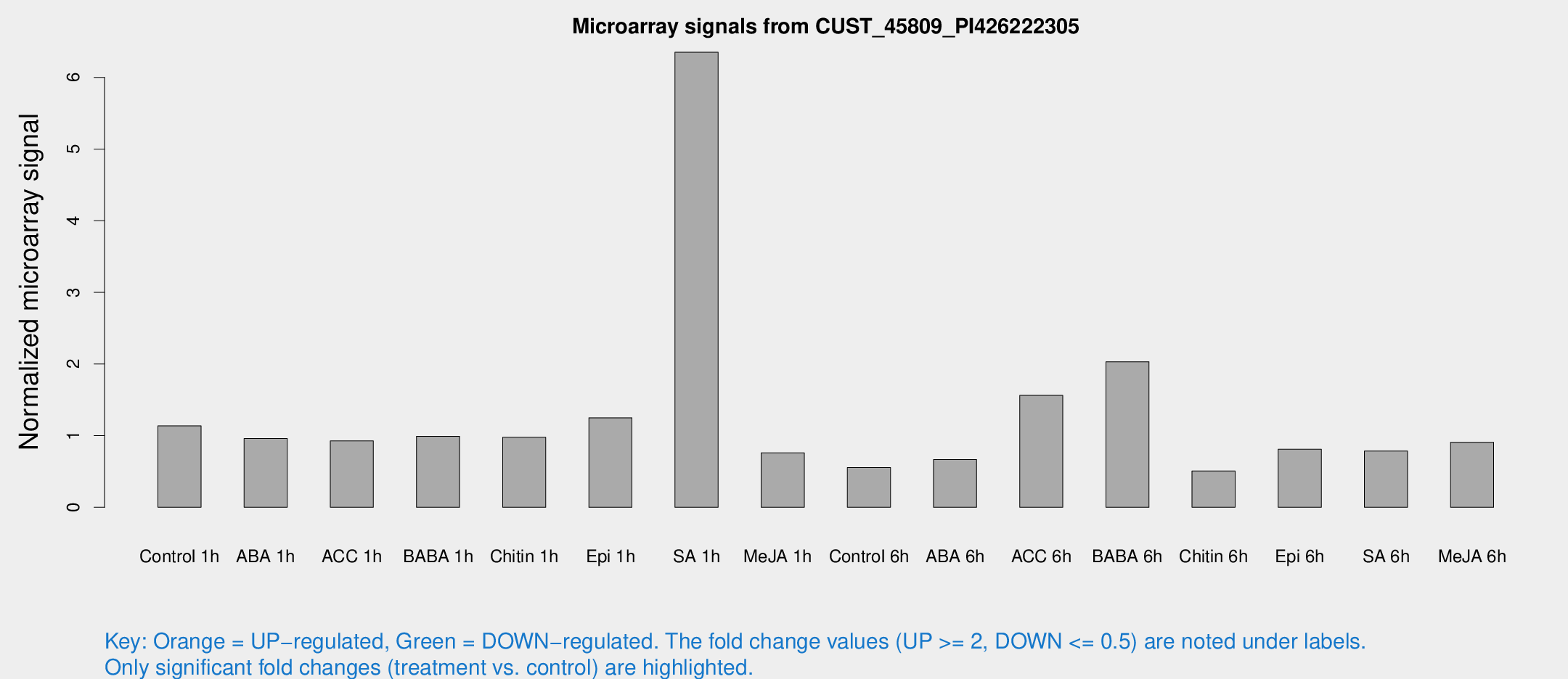
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 17.8473 | 5.72349 | 1.13742 | 0.329619 |
| ABA 1h | 13.5809 | 4.64331 | 0.959475 | 0.343031 |
| ACC 1h | 14.6615 | 4.68294 | 0.927255 | 0.365464 |
| BABA 1h | 15.1064 | 4.36966 | 0.991444 | 0.347457 |
| Chitin 1h | 12.9507 | 3.68521 | 0.977164 | 0.30439 |
| Epi 1h | 15.8558 | 3.73341 | 1.24969 | 0.327881 |
| SA 1h | 98.1667 | 23.2335 | 6.35231 | 1.43214 |
| Me-JA 1h | 9.8444 | 3.78515 | 0.75974 | 0.359723 |
| Control 6h | 8.09298 | 3.72866 | 0.555548 | 0.276274 |
| ABA 6h | 11.7891 | 5.08687 | 0.666563 | 0.305385 |
| ACC 6h | 25.8822 | 4.87231 | 1.56171 | 0.35083 |
| BABA 6h | 32.6772 | 5.09771 | 2.03184 | 0.288722 |
| Chitin 6h | 7.62715 | 4.09109 | 0.506459 | 0.274944 |
| Epi 6h | 16.7604 | 8.8405 | 0.810624 | 0.472859 |
| SA 6h | 11.3837 | 4.00744 | 0.785159 | 0.311051 |
| Me-JA 6h | 14.9732 | 5.78639 | 0.907997 | 0.331961 |
Source Transcript PGSC0003DMT400050269 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G14250.1 | +3 | 2e-56 | 191 | 98/305 (32%) | RING/U-box superfamily protein | chr3:4745963-4746958 REVERSE LENGTH=303 |
