Probe CUST_45376_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45376_PI426222305 | JHI_St_60k_v1 | DMT400001613 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
All Microarray Probes Designed to Gene DMG400000597
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45346_PI426222305 | JHI_St_60k_v1 | DMT400001614 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
| CUST_45376_PI426222305 | JHI_St_60k_v1 | DMT400001613 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
Microarray Signals from CUST_45376_PI426222305
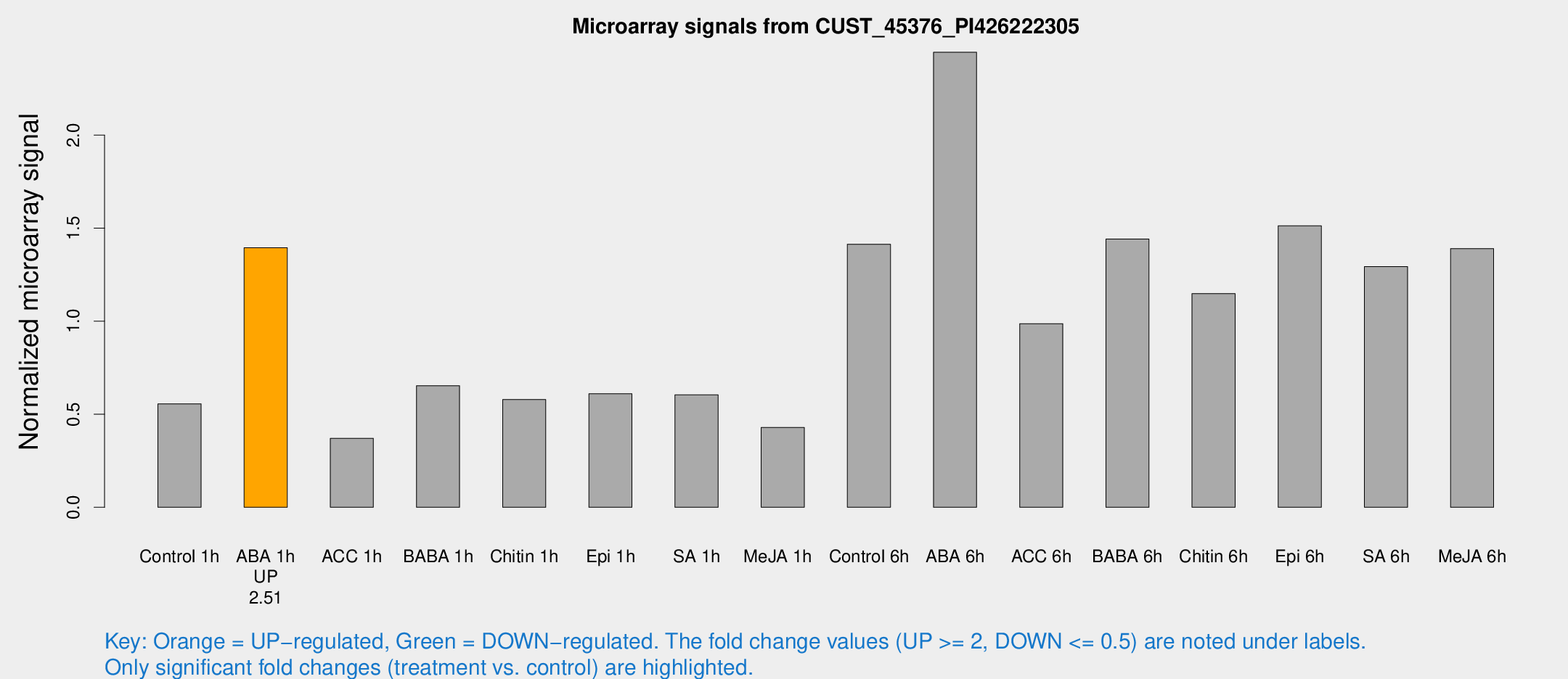
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 202.559 | 41.0009 | 0.556228 | 0.107823 |
| ABA 1h | 438.014 | 57.0209 | 1.39528 | 0.0866839 |
| ACC 1h | 140.779 | 32.4441 | 0.370681 | 0.0690076 |
| BABA 1h | 228.824 | 43.2818 | 0.6534 | 0.0705361 |
| Chitin 1h | 182.784 | 13.7698 | 0.579269 | 0.0353094 |
| Epi 1h | 202.872 | 54.7638 | 0.609939 | 0.206382 |
| SA 1h | 219.734 | 26.254 | 0.604375 | 0.0658952 |
| Me-JA 1h | 125.332 | 20.5227 | 0.429308 | 0.0375467 |
| Control 6h | 515.319 | 97.9377 | 1.41381 | 0.163714 |
| ABA 6h | 941.096 | 185.163 | 2.44543 | 0.334543 |
| ACC 6h | 403.222 | 55.3187 | 0.98669 | 0.141979 |
| BABA 6h | 569.617 | 63.0197 | 1.44152 | 0.104664 |
| Chitin 6h | 426.545 | 25.0045 | 1.14836 | 0.0672982 |
| Epi 6h | 602.84 | 71.1612 | 1.51272 | 0.200836 |
| SA 6h | 464.604 | 85.4808 | 1.29322 | 0.11544 |
| Me-JA 6h | 519.69 | 123.765 | 1.39043 | 0.302798 |
Source Transcript PGSC0003DMT400001613 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G47620.1 | +2 | 3e-45 | 168 | 155/312 (50%) | TEOSINTE BRANCHED, cycloidea and PCF (TCP) 14 | chr3:17559004-17560473 FORWARD LENGTH=489 |
