Probe CUST_45346_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45346_PI426222305 | JHI_St_60k_v1 | DMT400001614 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
All Microarray Probes Designed to Gene DMG400000597
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45346_PI426222305 | JHI_St_60k_v1 | DMT400001614 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
| CUST_45376_PI426222305 | JHI_St_60k_v1 | DMT400001613 | GGCCACAGTTTTAACAGAAGAAGGAGTATTTGATGAGTGATTTTGTCAAAATTTCTCTGA |
Microarray Signals from CUST_45346_PI426222305
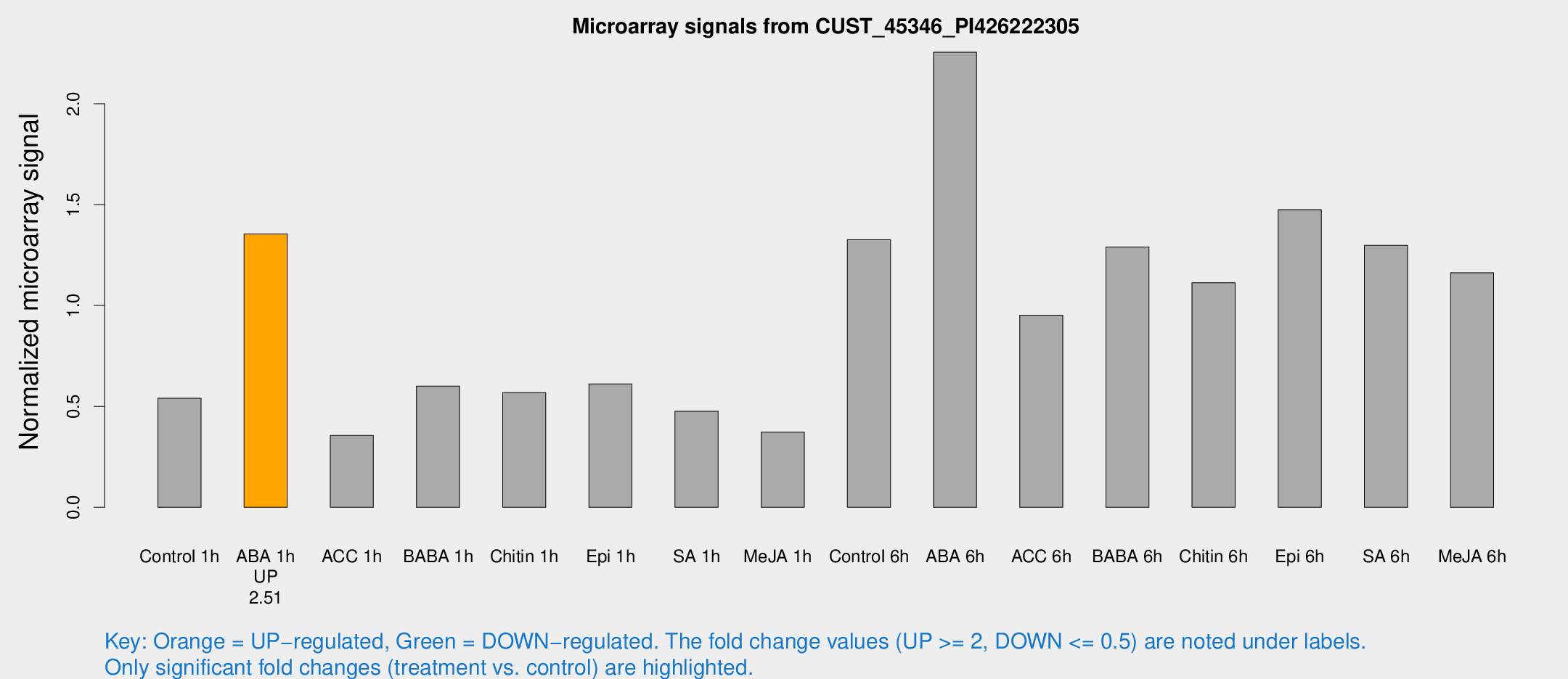
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 208.241 | 25.9253 | 0.54054 | 0.0480955 |
| ABA 1h | 463.421 | 63.9477 | 1.35465 | 0.0902677 |
| ACC 1h | 144.902 | 27.6418 | 0.356163 | 0.0503912 |
| BABA 1h | 230.911 | 47.6562 | 0.600606 | 0.0782411 |
| Chitin 1h | 195.046 | 14.533 | 0.568339 | 0.0340649 |
| Epi 1h | 214.01 | 49.5047 | 0.611253 | 0.155117 |
| SA 1h | 187.155 | 16.8623 | 0.475885 | 0.050079 |
| Me-JA 1h | 117.088 | 14.2167 | 0.372613 | 0.0287506 |
| Control 6h | 531.483 | 111.856 | 1.32581 | 0.190603 |
| ABA 6h | 945.832 | 188.501 | 2.25486 | 0.320153 |
| ACC 6h | 416.368 | 26.2697 | 0.952272 | 0.0978253 |
| BABA 6h | 552.298 | 48.5694 | 1.28989 | 0.108463 |
| Chitin 6h | 450.725 | 26.3546 | 1.11266 | 0.0649065 |
| Epi 6h | 636.585 | 60.303 | 1.47486 | 0.164607 |
| SA 6h | 501.106 | 78.5529 | 1.29748 | 0.0755707 |
| Me-JA 6h | 473.279 | 115.006 | 1.16262 | 0.264604 |
Source Transcript PGSC0003DMT400001614 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G47620.1 | +1 | 9e-43 | 161 | 152/309 (49%) | TEOSINTE BRANCHED, cycloidea and PCF (TCP) 14 | chr3:17559004-17560473 FORWARD LENGTH=489 |
