Probe CUST_45322_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45322_PI426222305 | JHI_St_60k_v1 | DMT400001687 | GTTAGAGTAGAATGGAATCGAATACATCTTGTAGAATTGAACACTCTGTAAAGTTTCACC |
All Microarray Probes Designed to Gene DMG400000631
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45322_PI426222305 | JHI_St_60k_v1 | DMT400001687 | GTTAGAGTAGAATGGAATCGAATACATCTTGTAGAATTGAACACTCTGTAAAGTTTCACC |
| CUST_45329_PI426222305 | JHI_St_60k_v1 | DMT400001688 | TTTCCAGTGTGAGAGCATAGATGCAGTGGAGAAGAAGCTGACAGAAATGGGGATAAAATA |
Microarray Signals from CUST_45322_PI426222305
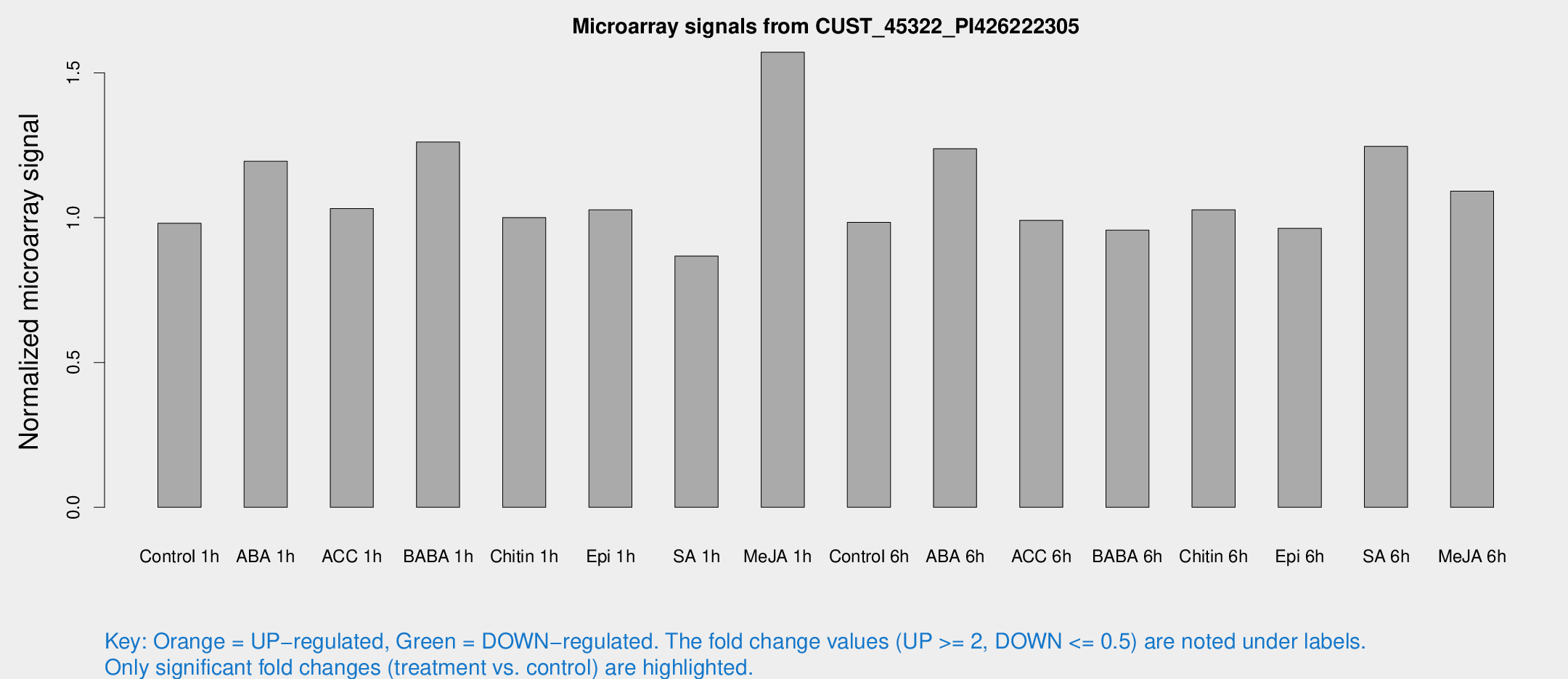
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.19419 | 3.34012 | 0.980785 | 0.529768 |
| ABA 1h | 6.99481 | 3.13398 | 1.1948 | 0.589883 |
| ACC 1h | 6.74473 | 3.94952 | 1.0314 | 0.597313 |
| BABA 1h | 7.88636 | 3.7094 | 1.26118 | 0.630739 |
| Chitin 1h | 5.69555 | 3.31402 | 1.00058 | 0.579424 |
| Epi 1h | 5.61663 | 3.26133 | 1.02711 | 0.59477 |
| SA 1h | 5.6297 | 3.27003 | 0.867273 | 0.502224 |
| Me-JA 1h | 8.91609 | 3.46554 | 1.57144 | 0.740438 |
| Control 6h | 6.22207 | 3.6093 | 0.983783 | 0.569759 |
| ABA 6h | 8.76007 | 3.77919 | 1.23816 | 0.601646 |
| ACC 6h | 7.3137 | 4.33928 | 0.990683 | 0.574479 |
| BABA 6h | 6.77183 | 3.93513 | 0.956919 | 0.554134 |
| Chitin 6h | 6.90391 | 4.00472 | 1.02753 | 0.594965 |
| Epi 6h | 6.93454 | 4.07025 | 0.963257 | 0.55778 |
| SA 6h | 7.97261 | 3.79764 | 1.24649 | 0.628523 |
| Me-JA 6h | 6.91337 | 3.56138 | 1.09172 | 0.573993 |
Source Transcript PGSC0003DMT400001687 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G80160.1 | +3 | 8e-41 | 145 | 66/87 (76%) | Lactoylglutathione lyase / glyoxalase I family protein | chr1:30151101-30151930 FORWARD LENGTH=167 |
