Probe CUST_45192_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45192_PI426222305 | JHI_St_60k_v1 | DMT400081222 | TTGCTTGTTAGTTGTCGAATTCTTGTTTAGCCGATCCCTCTTATATCTCGTATAACTGTT |
All Microarray Probes Designed to Gene DMG400031750
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45145_PI426222305 | JHI_St_60k_v1 | DMT400081223 | ATCGACTCCTCATGTACTTTGCAGAGGGAGGTGGATTATTTATGTGGAGCACAAACTTGC |
| CUST_45192_PI426222305 | JHI_St_60k_v1 | DMT400081222 | TTGCTTGTTAGTTGTCGAATTCTTGTTTAGCCGATCCCTCTTATATCTCGTATAACTGTT |
Microarray Signals from CUST_45192_PI426222305
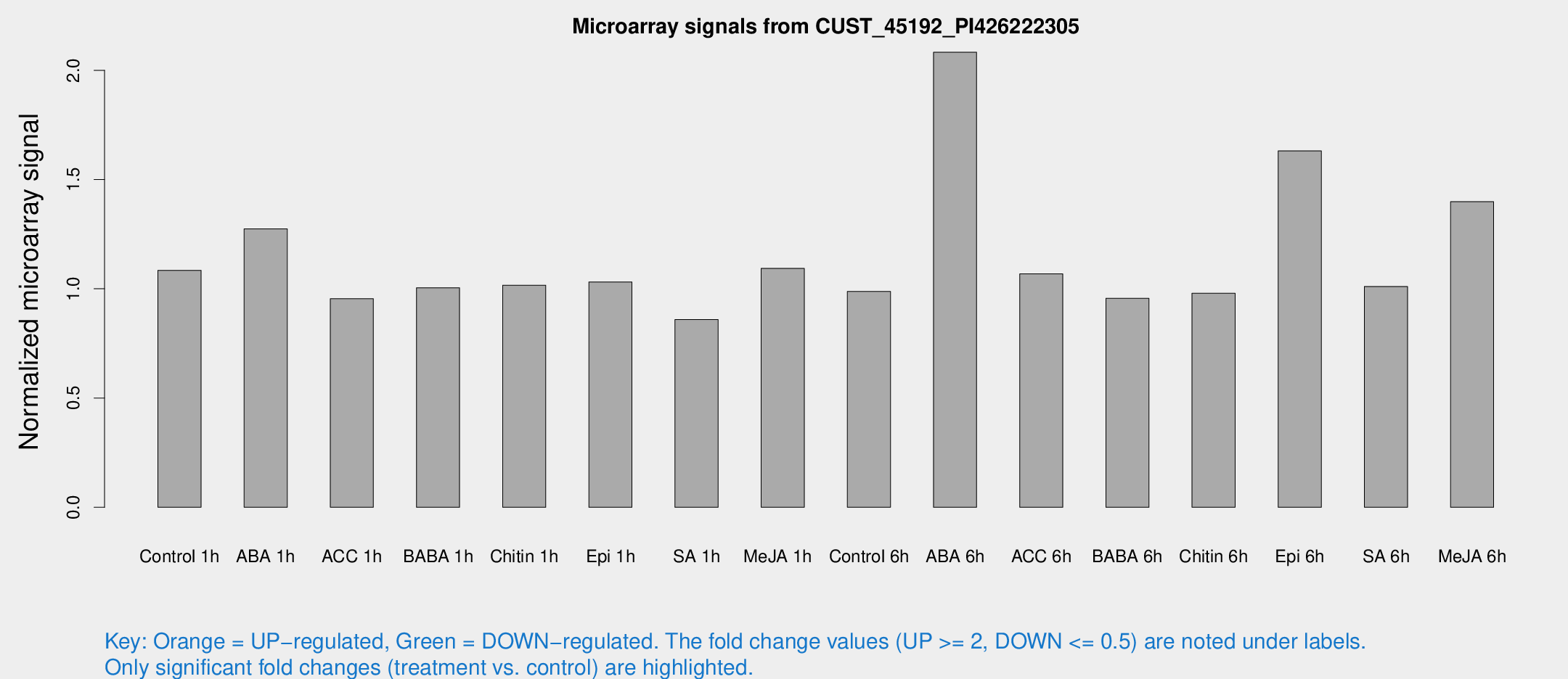
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.90679 | 3.64467 | 1.08459 | 0.540424 |
| ABA 1h | 8.41496 | 3.54733 | 1.27392 | 0.614208 |
| ACC 1h | 6.90124 | 4.00637 | 0.954344 | 0.552745 |
| BABA 1h | 6.80462 | 3.94676 | 1.00456 | 0.582512 |
| Chitin 1h | 6.4356 | 3.73468 | 1.01655 | 0.588672 |
| Epi 1h | 6.28015 | 3.64179 | 1.03144 | 0.597276 |
| SA 1h | 6.20752 | 3.59988 | 0.85905 | 0.497541 |
| Me-JA 1h | 6.2944 | 3.65803 | 1.0936 | 0.633211 |
| Control 6h | 6.95471 | 3.92769 | 0.987518 | 0.557768 |
| ABA 6h | 15.5591 | 4.17405 | 2.08263 | 0.559908 |
| ACC 6h | 8.74178 | 4.76995 | 1.06837 | 0.570349 |
| BABA 6h | 7.55183 | 4.27493 | 0.956574 | 0.539596 |
| Chitin 6h | 7.32543 | 4.24382 | 0.979319 | 0.567197 |
| Epi 6h | 15.7251 | 7.26275 | 1.63109 | 0.983782 |
| SA 6h | 7.02312 | 4.07293 | 1.01067 | 0.586114 |
| Me-JA 6h | 11.915 | 5.52733 | 1.39839 | 0.64743 |
Source Transcript PGSC0003DMT400081222 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G20560.1 | +3 | 1e-74 | 237 | 145/205 (71%) | DNAJ heat shock family protein | chr2:8848353-8849815 REVERSE LENGTH=337 |
