Probe CUST_45011_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_45011_PI426222305 | JHI_St_60k_v1 | DMT400014068 | GACCTACTATTGTTGCTCAACAAGGTGACACTATTGTTGTTGAAGTCAAAAATAGTTTGT |
All Microarray Probes Designed to Gene DMG400005515
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44995_PI426222305 | JHI_St_60k_v1 | DMT400014067 | CTGTGGAGAAAGTAAAAGGTTCCTCAGGCCTTAAATTCTTTAACATTATTGTCCAAAAGA |
| CUST_45011_PI426222305 | JHI_St_60k_v1 | DMT400014068 | GACCTACTATTGTTGCTCAACAAGGTGACACTATTGTTGTTGAAGTCAAAAATAGTTTGT |
Microarray Signals from CUST_45011_PI426222305
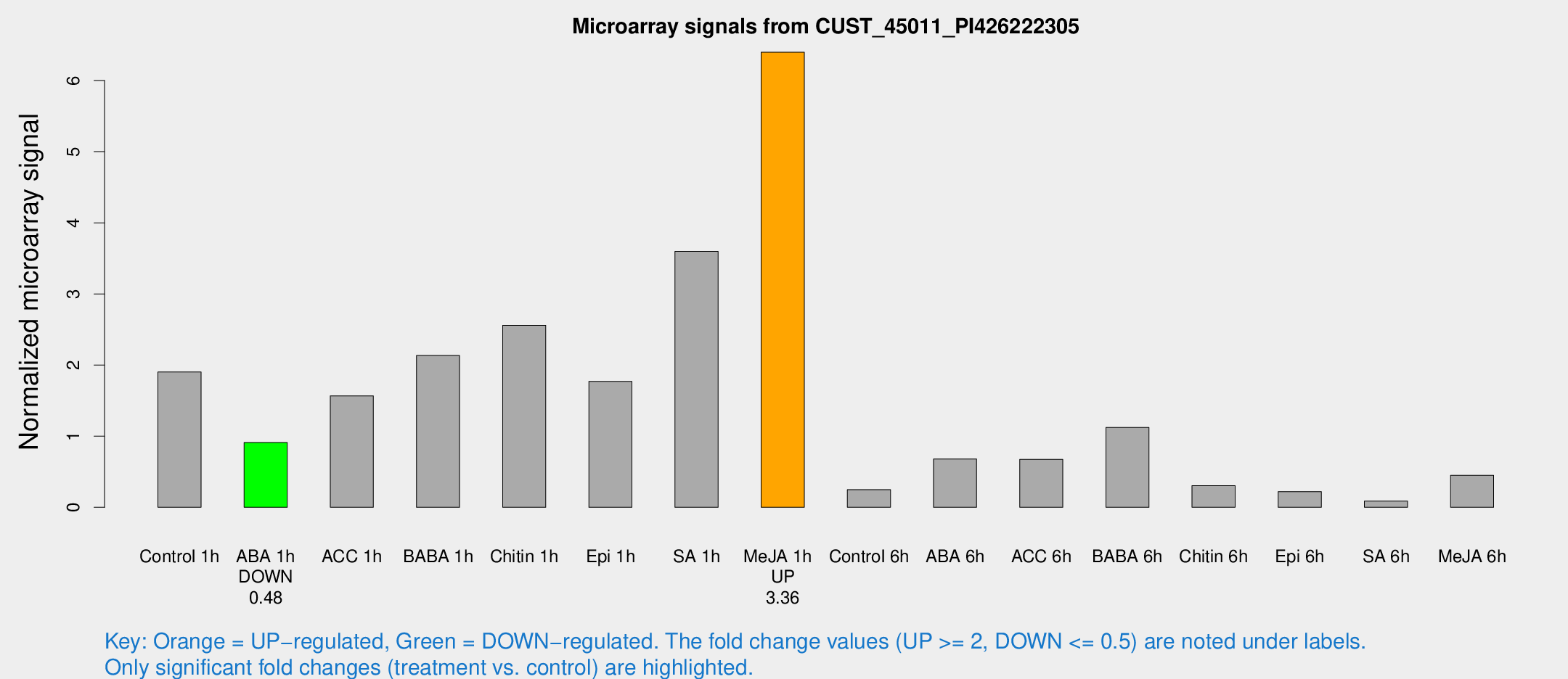
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 224.153 | 55.8061 | 1.90196 | 0.35593 |
| ABA 1h | 90.6459 | 9.81599 | 0.909799 | 0.147747 |
| ACC 1h | 262.265 | 113.22 | 1.56748 | 1.44022 |
| BABA 1h | 250.234 | 65.6067 | 2.13457 | 0.462539 |
| Chitin 1h | 256.344 | 15.2111 | 2.55869 | 0.214178 |
| Epi 1h | 170.949 | 10.4606 | 1.77103 | 0.111292 |
| SA 1h | 444.84 | 109.024 | 3.59854 | 0.898205 |
| Me-JA 1h | 594.999 | 87.0104 | 6.39807 | 0.435115 |
| Control 6h | 42.3137 | 20.0653 | 0.248859 | 0.23617 |
| ABA 6h | 103.092 | 39.9652 | 0.679331 | 0.426084 |
| ACC 6h | 88.6619 | 15.1861 | 0.674066 | 0.0662582 |
| BABA 6h | 165.904 | 68.7431 | 1.12323 | 0.463303 |
| Chitin 6h | 37.8341 | 7.7285 | 0.304056 | 0.069945 |
| Epi 6h | 33.7914 | 13.889 | 0.220889 | 0.105399 |
| SA 6h | 9.99235 | 3.85469 | 0.0871447 | 0.0371988 |
| Me-JA 6h | 58.7354 | 21.688 | 0.44909 | 0.167401 |
Source Transcript PGSC0003DMT400014068 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G39830.1 | +2 | 0.0 | 809 | 391/573 (68%) | Cupredoxin superfamily protein | chr4:18479103-18481184 FORWARD LENGTH=582 |
