Probe CUST_44995_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44995_PI426222305 | JHI_St_60k_v1 | DMT400014067 | CTGTGGAGAAAGTAAAAGGTTCCTCAGGCCTTAAATTCTTTAACATTATTGTCCAAAAGA |
All Microarray Probes Designed to Gene DMG400005515
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44995_PI426222305 | JHI_St_60k_v1 | DMT400014067 | CTGTGGAGAAAGTAAAAGGTTCCTCAGGCCTTAAATTCTTTAACATTATTGTCCAAAAGA |
| CUST_45011_PI426222305 | JHI_St_60k_v1 | DMT400014068 | GACCTACTATTGTTGCTCAACAAGGTGACACTATTGTTGTTGAAGTCAAAAATAGTTTGT |
Microarray Signals from CUST_44995_PI426222305
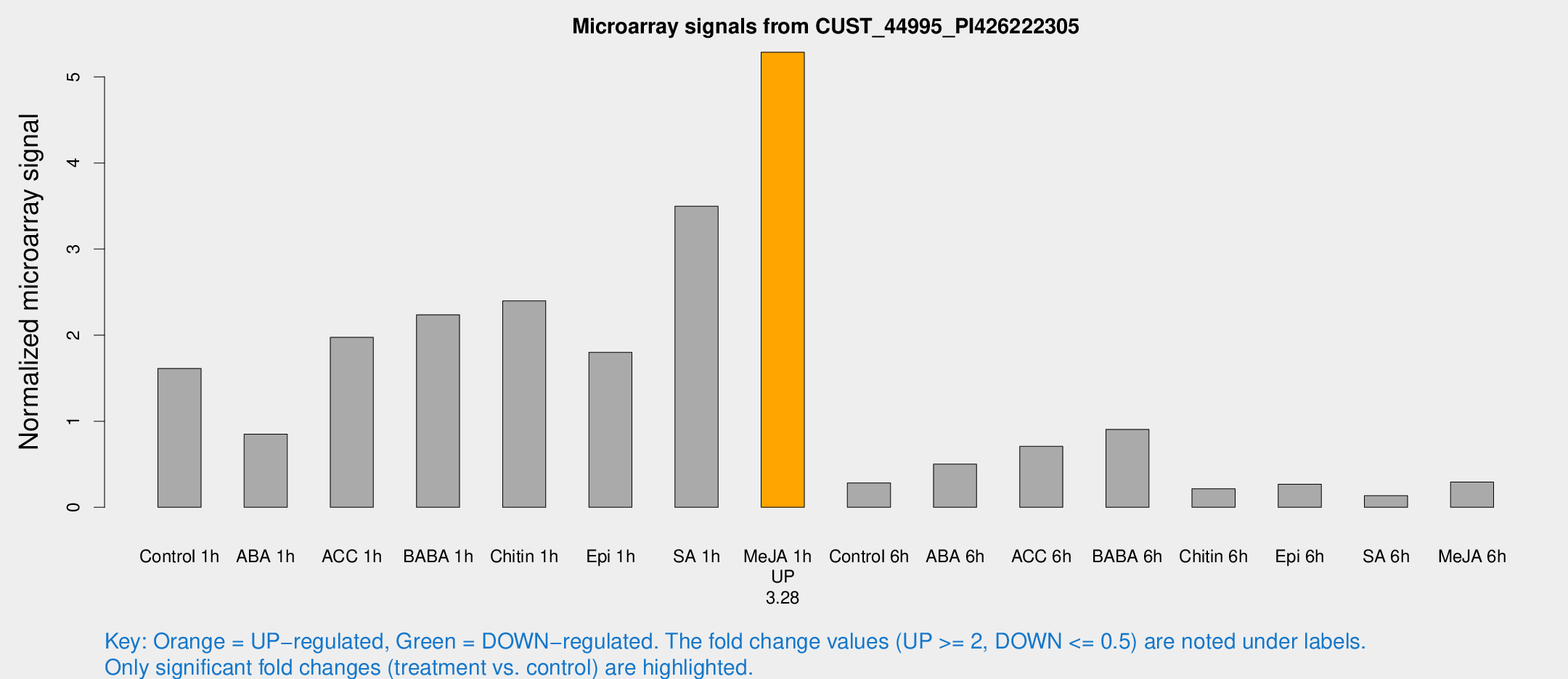
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8288.45 | 1705.97 | 1.61258 | 0.249269 |
| ABA 1h | 3834.93 | 643.455 | 0.850508 | 0.182659 |
| ACC 1h | 11165.4 | 3390.36 | 1.97483 | 0.572835 |
| BABA 1h | 11418.6 | 2707.11 | 2.23629 | 0.395147 |
| Chitin 1h | 10672.7 | 617.057 | 2.39903 | 0.282646 |
| Epi 1h | 7710.09 | 445.917 | 1.79988 | 0.119923 |
| SA 1h | 18912.8 | 4574.64 | 3.49822 | 0.815101 |
| Me-JA 1h | 21607.6 | 2544.09 | 5.28675 | 0.305232 |
| Control 6h | 1635.43 | 605.341 | 0.283646 | 0.0949765 |
| ABA 6h | 2976.7 | 1008.87 | 0.502034 | 0.159568 |
| ACC 6h | 4257.52 | 934.872 | 0.708483 | 0.195384 |
| BABA 6h | 5593.42 | 1946.26 | 0.905312 | 0.290186 |
| Chitin 6h | 1152.49 | 143.735 | 0.215586 | 0.0262092 |
| Epi 6h | 1641.65 | 429.202 | 0.26886 | 0.0723402 |
| SA 6h | 665.714 | 54.0918 | 0.135269 | 0.025698 |
| Me-JA 6h | 1576.04 | 484.82 | 0.293977 | 0.070195 |
Source Transcript PGSC0003DMT400014067 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G39830.1 | +2 | 1e-157 | 460 | 226/323 (70%) | Cupredoxin superfamily protein | chr4:18479103-18481184 FORWARD LENGTH=582 |
