Probe CUST_44932_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44932_PI426222305 | JHI_St_60k_v1 | DMT400058675 | CTATGGATGATCCAAGCTCAAGGTTACGGGAAATTTTAAGTATGAATGTTGATTGTATTG |
All Microarray Probes Designed to Gene DMG400022794
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44932_PI426222305 | JHI_St_60k_v1 | DMT400058675 | CTATGGATGATCCAAGCTCAAGGTTACGGGAAATTTTAAGTATGAATGTTGATTGTATTG |
Microarray Signals from CUST_44932_PI426222305
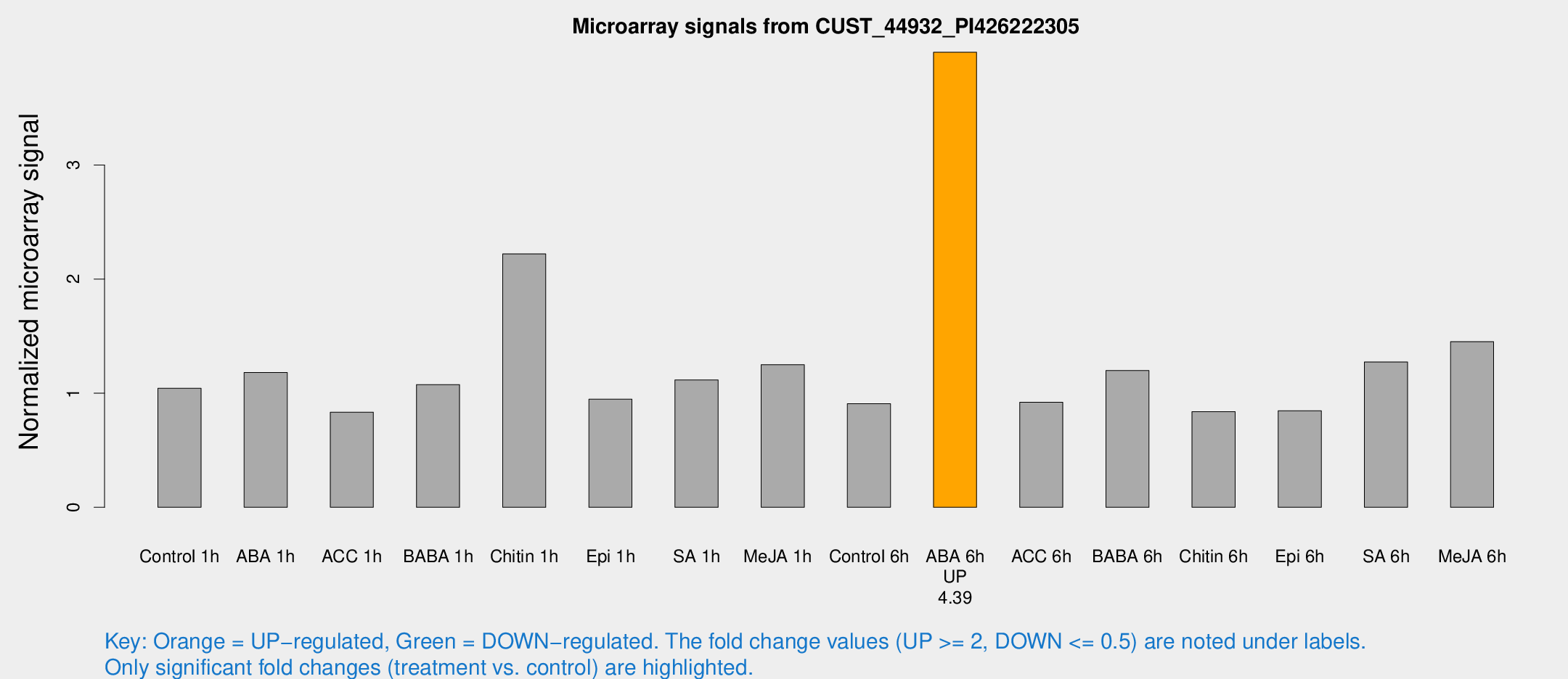
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.6333 | 3.45103 | 1.04345 | 0.496876 |
| ABA 1h | 8.12981 | 3.38578 | 1.18122 | 0.520328 |
| ACC 1h | 6.53686 | 3.79865 | 0.833521 | 0.483033 |
| BABA 1h | 8.17596 | 3.69607 | 1.07556 | 0.520537 |
| Chitin 1h | 18.4366 | 7.69187 | 2.22025 | 0.917522 |
| Epi 1h | 6.25272 | 3.47903 | 0.947211 | 0.527248 |
| SA 1h | 10.2105 | 4.23721 | 1.11631 | 0.518698 |
| Me-JA 1h | 7.98476 | 3.56672 | 1.2504 | 0.582314 |
| Control 6h | 7.01407 | 3.65366 | 0.90767 | 0.483114 |
| ABA 6h | 33.29 | 5.49757 | 3.98849 | 0.594892 |
| ACC 6h | 8.18503 | 4.47316 | 0.920576 | 0.488491 |
| BABA 6h | 11.3484 | 4.11269 | 1.19839 | 0.547936 |
| Chitin 6h | 6.8018 | 3.94005 | 0.838948 | 0.485832 |
| Epi 6h | 7.34457 | 4.21333 | 0.846685 | 0.483321 |
| SA 6h | 11.4896 | 5.11902 | 1.27322 | 0.844099 |
| Me-JA 6h | 12.1477 | 3.75445 | 1.45121 | 0.550847 |
Source Transcript PGSC0003DMT400058675 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G52770.1 | +2 | 3e-08 | 48 | 23/48 (48%) | protein binding | chr3:19557891-19558094 REVERSE LENGTH=67 |
