Probe CUST_44882_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44882_PI426222305 | JHI_St_60k_v1 | DMT400044097 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
All Microarray Probes Designed to Gene DMG400017110
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_44879_PI426222305 | JHI_St_60k_v1 | DMT400044098 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
| CUST_44882_PI426222305 | JHI_St_60k_v1 | DMT400044097 | GTATGGGATACCAACATCAAGTATTCTCAAGGCAGGGGCATTGGCATTGTTGTTAGTCCA |
Microarray Signals from CUST_44882_PI426222305
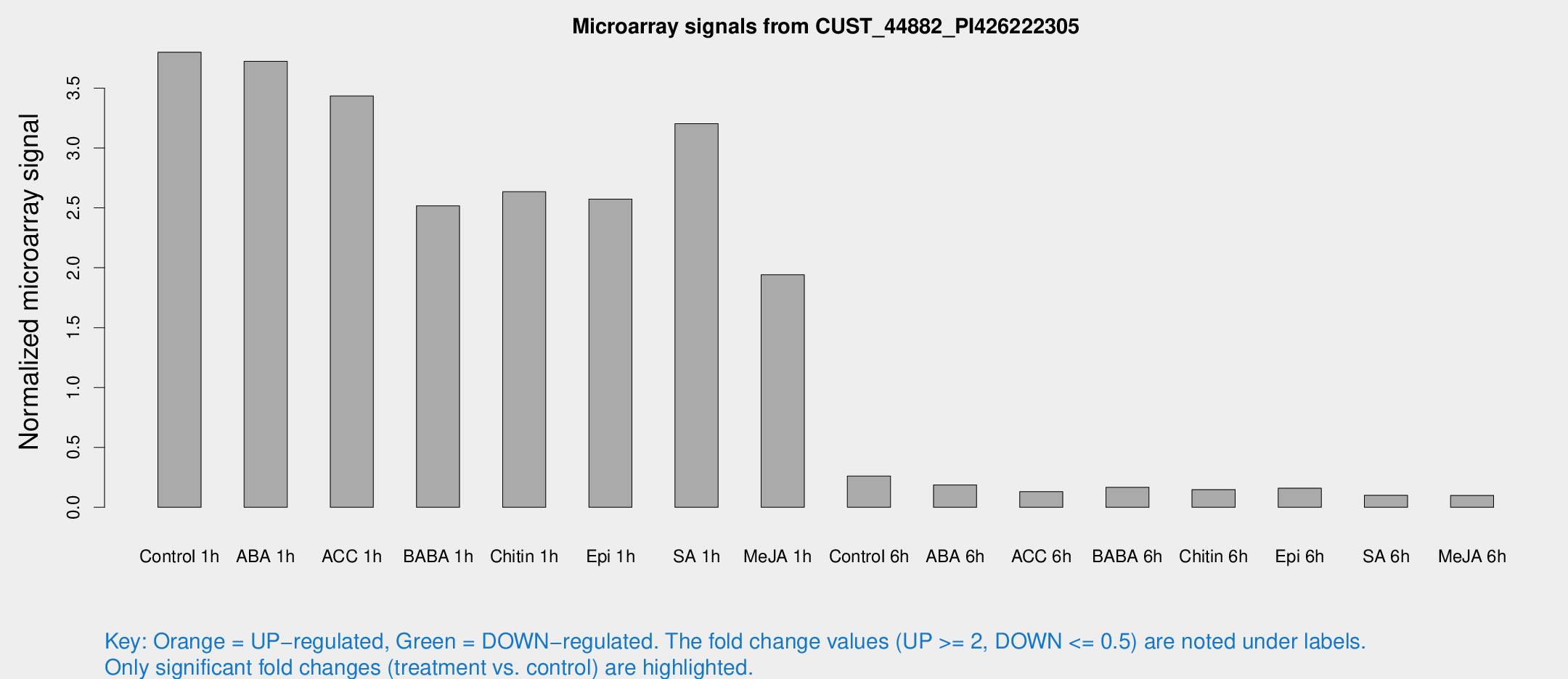
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10100.4 | 1277.88 | 3.79907 | 0.222547 |
| ABA 1h | 8632.71 | 499.068 | 3.72311 | 0.276042 |
| ACC 1h | 9348.89 | 1011.08 | 3.43347 | 0.198237 |
| BABA 1h | 6919.7 | 1765.33 | 2.51645 | 0.534871 |
| Chitin 1h | 6300.5 | 790.704 | 2.63429 | 0.291356 |
| Epi 1h | 5832.25 | 336.866 | 2.57324 | 0.148574 |
| SA 1h | 8685.41 | 830.373 | 3.20185 | 0.184864 |
| Me-JA 1h | 4203.79 | 483.971 | 1.94074 | 0.112062 |
| Control 6h | 725.332 | 162.661 | 0.260767 | 0.0411744 |
| ABA 6h | 525.754 | 57.2484 | 0.186718 | 0.0159056 |
| ACC 6h | 402.154 | 59.3244 | 0.131232 | 0.00771151 |
| BABA 6h | 491.552 | 44.4525 | 0.166281 | 0.0216606 |
| Chitin 6h | 421.415 | 62.6524 | 0.148011 | 0.0243742 |
| Epi 6h | 501.743 | 130.717 | 0.159596 | 0.0448496 |
| SA 6h | 278.08 | 62.6577 | 0.100766 | 0.0150152 |
| Me-JA 6h | 279.526 | 67.1799 | 0.0996389 | 0.0211894 |
Source Transcript PGSC0003DMT400044097 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G32530.1 | +1 | 0.0 | 611 | 356/766 (46%) | cellulose synthase-like B3 | chr2:13809283-13813487 FORWARD LENGTH=755 |
